Genomic screening with RNAi: results and challenges
- PMID: 20367032
- PMCID: PMC3564595
- DOI: 10.1146/annurev-biochem-060408-092949
Genomic screening with RNAi: results and challenges
Abstract
RNA interference (RNAi) is an effective tool for genome-scale, high-throughput analysis of gene function. In the past five years, a number of genome-scale RNAi high-throughput screens (HTSs) have been done in both Drosophila and mammalian cultured cells to study diverse biological processes, including signal transduction, cancer biology, and host cell responses to infection. Results from these screens have led to the identification of new components of these processes and, importantly, have also provided insights into the complexity of biological systems, forcing new and innovative approaches to understanding functional networks in cells. Here, we review the main findings that have emerged from RNAi HTS and discuss technical issues that remain to be improved, in particular the verification of RNAi results and validation of their biological relevance. Furthermore, we discuss the importance of multiplexed and integrated experimental data analysis pipelines to RNAi HTS.
Figures
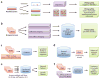
References
-
- Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 1998;391:806–11. - PubMed
-
- Zamore PD, Tuschl T, Sharp PA, Bartel DP. RNAi: double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals. Cell. 2000;101:25–33. - PubMed
-
- Kim VN, Han J, Siomi MC. Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol. 2009;10:126–39. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous

