A global in vivo Drosophila RNAi screen identifies NOT3 as a conserved regulator of heart function
- PMID: 20371351
- PMCID: PMC2855221
- DOI: 10.1016/j.cell.2010.02.023
A global in vivo Drosophila RNAi screen identifies NOT3 as a conserved regulator of heart function
Abstract
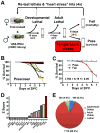
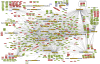
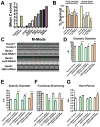
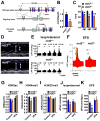
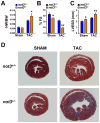
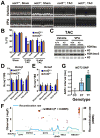
Heart diseases are the most common causes of morbidity and death in humans. Using cardiac-specific RNAi-silencing in Drosophila, we knocked down 7061 evolutionarily conserved genes under conditions of stress. We present a first global roadmap of pathways potentially playing conserved roles in the cardiovascular system. One critical pathway identified was the CCR4-Not complex implicated in transcriptional and posttranscriptional regulatory mechanisms. Silencing of CCR4-Not components in adult Drosophila resulted in myofibrillar disarray and dilated cardiomyopathy. Heterozygous not3 knockout mice showed spontaneous impairment of cardiac contractility and increased susceptibility to heart failure. These heart defects were reversed via inhibition of HDACs, suggesting a mechanistic link to epigenetic chromatin remodeling. In humans, we show that a common NOT3 SNP correlates with altered cardiac QT intervals, a known cause of potentially lethal ventricular tachyarrhythmias. Thus, our functional genome-wide screen in Drosophila can identify candidates that directly translate into conserved mammalian genes involved in heart function.
Copyright 2010 Elsevier Inc. All rights reserved.
Figures

References
-
- A.H.A. 2005. http://www.americanheart.org/
-
- Bodmer R. Heart development in Drosophila and its relationship to vertebrates Trends in cardiovascular medicine. 1995;5:21–28. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

