Redox sensing by a Rex-family repressor is involved in the regulation of anaerobic gene expression in Staphylococcus aureus
- PMID: 20374494
- PMCID: PMC2883068
- DOI: 10.1111/j.1365-2958.2010.07105.x
Redox sensing by a Rex-family repressor is involved in the regulation of anaerobic gene expression in Staphylococcus aureus
Abstract
An alignment of upstream regions of anaerobically induced genes in Staphylococcus aureus revealed the presence of an inverted repeat, corresponding to Rex binding sites in Streptomyces coelicolor. Gel shift experiments of selected upstream regions demonstrated that the redox-sensing regulator Rex of S. aureus binds to this inverted repeat. The binding sequence--TTGTGAAW(4)TTCACAA--is highly conserved in S. aureus. Rex binding to this sequence leads to the repression of genes located downstream. The binding activity of Rex is enhanced by NAD+ while NADH, which competes with NAD+ for Rex binding, decreases the activity of Rex. The impact of Rex on global protein synthesis and on the activity of fermentation pathways under aerobic and anaerobic conditions was analysed by using a rex-deficient strain. A direct regulatory effect of Rex on the expression of pathways that lead to anaerobic NAD+ regeneration, such as lactate, formate and ethanol formation, nitrate respiration, and ATP synthesis, is verified. Rex can be considered a central regulator of anaerobic metabolism in S. aureus. Since the activity of lactate dehydrogenase enables S. aureus to resist NO stress and thus the innate immune response, our data suggest that deactivation of Rex is a prerequisite for this phenomenon.
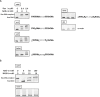
Figures








Similar articles
-
A novel sensor of NADH/NAD+ redox poise in Streptomyces coelicolor A3(2).EMBO J. 2003 Sep 15;22(18):4856-65. doi: 10.1093/emboj/cdg453. EMBO J. 2003. PMID: 12970197 Free PMC article.
-
Regulatory loop between redox sensing of the NADH/NAD(+) ratio by Rex (YdiH) and oxidation of NADH by NADH dehydrogenase Ndh in Bacillus subtilis.J Bacteriol. 2006 Oct;188(20):7062-71. doi: 10.1128/JB.00601-06. J Bacteriol. 2006. PMID: 17015645 Free PMC article.
-
The Intersection of the Staphylococcus aureus Rex and SrrAB Regulons: an Example of Metabolic Evolution That Maximizes Resistance to Immune Radicals.mBio. 2021 Dec 21;12(6):e0218821. doi: 10.1128/mBio.02188-21. Epub 2021 Nov 16. mBio. 2021. PMID: 34781744 Free PMC article.
-
[Cloning and expression of the redox-sensing transcriptional repressor Rex and in vitro DNA-binding assay of the Rex and rex operator in Streptomyces rimosus M4018].Wei Sheng Wu Xue Bao. 2012 Jan;52(1):38-43. Wei Sheng Wu Xue Bao. 2012. PMID: 22489458 Chinese.
-
Transcription factor Rex in regulation of pathophysiology in oral pathogens.Mol Oral Microbiol. 2016 Apr;31(2):115-24. doi: 10.1111/omi.12114. Epub 2015 Aug 6. Mol Oral Microbiol. 2016. PMID: 26172563 Free PMC article. Review.
Cited by
-
Streptococcus pneumoniae metal homeostasis alters cellular metabolism.Metallomics. 2020 Sep 23;12(9):1416-1427. doi: 10.1039/d0mt00118j. Metallomics. 2020. PMID: 32676626 Free PMC article.
-
RpiRc Is a Pleiotropic Effector of Virulence Determinant Synthesis and Attenuates Pathogenicity in Staphylococcus aureus.Infect Immun. 2016 Jun 23;84(7):2031-2041. doi: 10.1128/IAI.00285-16. Print 2016 Jul. Infect Immun. 2016. PMID: 27113358 Free PMC article.
-
Rex (encoded by DVU_0916) in Desulfovibrio vulgaris Hildenborough is a repressor of sulfate adenylyl transferase and is regulated by NADH.J Bacteriol. 2015 Jan 1;197(1):29-39. doi: 10.1128/JB.02083-14. Epub 2014 Oct 13. J Bacteriol. 2015. PMID: 25313388 Free PMC article.
-
Mutations in the Staphylococcus aureus Global Regulator CodY confer tolerance to an interspecies redox-active antimicrobial.PLoS Genet. 2025 Mar 7;21(3):e1011610. doi: 10.1371/journal.pgen.1011610. eCollection 2025 Mar. PLoS Genet. 2025. PMID: 40053555 Free PMC article.
-
An rpsL-based allelic exchange vector for Staphylococcus aureus.Plasmid. 2015 May;79:8-14. doi: 10.1016/j.plasmid.2015.02.002. Epub 2015 Feb 7. Plasmid. 2015. PMID: 25659529 Free PMC article.
References
-
- Bernhardt J, Büttner K, Scharf C, Hecker M. Dual channel imaging of two-dimensional electropherograms in Bacillus subtilis. Electrophoresis. 1999;20:2225–2240. - PubMed
-
- Brown GC, McBride AG, Fox EJ, McNaught KS, Borutaite V. Nitric oxide and oxygen metabolism. Biochem Soc Trans. 1997;25:901–904. - PubMed
-
- Büttner K, Bernhardt J, Scharf C, Schmid R, Mäder U, Eymann C, et al. A comprehensive two-dimensional map of cytosolic proteins of Bacillus subtilis. Electrophoresis. 2001;22:2908–2935. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

