Crystal structure of aminomethyltransferase in complex with dihydrolipoyl-H-protein of the glycine cleavage system: implications for recognition of lipoyl protein substrate, disease-related mutations, and reaction mechanism
- PMID: 20375021
- PMCID: PMC2881793
- DOI: 10.1074/jbc.M110.110718
Crystal structure of aminomethyltransferase in complex with dihydrolipoyl-H-protein of the glycine cleavage system: implications for recognition of lipoyl protein substrate, disease-related mutations, and reaction mechanism
Abstract
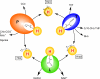
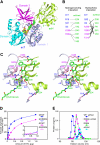
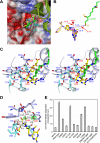
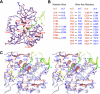
Aminomethyltransferase, a component of the glycine cleavage system termed T-protein, reversibly catalyzes the degradation of the aminomethyl moiety of glycine attached to the lipoate cofactor of H-protein, resulting in the production of ammonia, 5,10-methylenetetrahydrofolate, and dihydrolipoate-bearing H-protein in the presence of tetrahydrofolate. Several mutations in the human T-protein gene are known to cause nonketotic hyperglycinemia. Here, we report the crystal structure of Escherichia coli T-protein in complex with dihydrolipoate-bearing H-protein and 5-methyltetrahydrofolate, a complex mimicking the ternary complex in the reverse reaction. The structure of the complex shows a highly interacting intermolecular interface limited to a small area and the protein-bound dihydrolipoyllysine arm inserted into the active site cavity of the T-protein. Invariant Arg(292) of the T-protein is essential for complex assembly. The structure also provides novel insights in understanding the disease-causing mutations, in addition to the disease-related impairment in the cofactor-enzyme interactions reported previously. Furthermore, structural and mutational analyses suggest that the reversible transfer of the methylene group between the lipoate and tetrahydrofolate should proceed through the electron relay-assisted iminium intermediate formation.
Figures

Similar articles
-
Crystal structure of human T-protein of glycine cleavage system at 2.0 A resolution and its implication for understanding non-ketotic hyperglycinemia.J Mol Biol. 2005 Sep 2;351(5):1146-59. doi: 10.1016/j.jmb.2005.06.056. J Mol Biol. 2005. PMID: 16051266
-
Crystal structure of T-protein of the glycine cleavage system. Cofactor binding, insights into H-protein recognition, and molecular basis for understanding nonketotic hyperglycinemia.J Biol Chem. 2004 Nov 26;279(48):50514-23. doi: 10.1074/jbc.M409672200. Epub 2004 Sep 7. J Biol Chem. 2004. PMID: 15355973
-
Glycine cleavage system: reaction mechanism, physiological significance, and hyperglycinemia.Proc Jpn Acad Ser B Phys Biol Sci. 2008;84(7):246-63. doi: 10.2183/pjab.84.246. Proc Jpn Acad Ser B Phys Biol Sci. 2008. PMID: 18941301 Free PMC article. Review.
-
Mechanism of the glycine cleavage reaction. Properties of the reverse reaction catalyzed by T-protein.J Biol Chem. 1987 May 15;262(14):6746-9. J Biol Chem. 1987. PMID: 3571285
-
The ATP synthase glycine zipper of the c subunits: From the structural to the functional role in mitochondrial biology of cardiovascular diseases.Biochim Biophys Acta Mol Cell Res. 2021 Aug;1868(9):119075. doi: 10.1016/j.bbamcr.2021.119075. Epub 2021 Jun 8. Biochim Biophys Acta Mol Cell Res. 2021. PMID: 34111435 Review. No abstract available.
Cited by
-
Understanding and Engineering Glycine Cleavage System and Related Metabolic Pathways for C1-Based Biosynthesis.Adv Biochem Eng Biotechnol. 2022;180:273-298. doi: 10.1007/10_2021_186. Adv Biochem Eng Biotechnol. 2022. PMID: 35294558 Review.
-
VimR is a conserved master transcription regulator controlling virulence and metabolism in Pseudomonas.Nucleic Acids Res. 2025 Jun 6;53(11):gkaf493. doi: 10.1093/nar/gkaf493. Nucleic Acids Res. 2025. PMID: 40498071 Free PMC article.
-
Discovery of a sesamin-metabolizing microorganism and a new enzyme.Proc Natl Acad Sci U S A. 2016 Aug 9;113(32):9087-92. doi: 10.1073/pnas.1605050113. Epub 2016 Jul 21. Proc Natl Acad Sci U S A. 2016. PMID: 27444012 Free PMC article.
-
Proteomic analysis of the effect of retinoic acids on the human breast cancer cell line MCF-7.Mol Biol Rep. 2014 May;41(5):3499-507. doi: 10.1007/s11033-014-3212-8. Epub 2014 Mar 11. Mol Biol Rep. 2014. PMID: 24615502
-
Gene expression profiling gut microbiota in different races of humans.Sci Rep. 2016 Mar 15;6:23075. doi: 10.1038/srep23075. Sci Rep. 2016. PMID: 26975620 Free PMC article.
References
-
- Motokawa Y., Fujiwara K., Okamura-Ikeda K. (1995) in Biothiols in Health and Disease (Packer L., Cadenas E. eds) pp. 389–407, Marcel Dekker, New York
-
- Tada K., Hayasaka K. (1987) Eur. J. Pediatr. 146, 221–227 - PubMed
-
- Nanao K., Okamura-Ikeda K., Motokawa Y., Danks D. M., Baumgartner E. R., Takada G., Hayasaka K. (1994) Hum. Genet. 93, 655–658 - PubMed
-
- Okamura-Ikeda K., Hosaka H., Yoshimura M., Yamashita E., Toma S., Nakagawa A., Fujiwara K., Motokawa Y., Taniguchi H. (2005) J. Mol. Biol. 351, 1146–1159 - PubMed
-
- Okamura-Ikeda K., Fujiwara K., Motokawa Y. (1999) Eur. J. Biochem. 264, 446–452 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

