Flexibility between the protease and helicase domains of the dengue virus NS3 protein conferred by the linker region and its functional implications
- PMID: 20375022
- PMCID: PMC2881804
- DOI: 10.1074/jbc.M109.090936
Flexibility between the protease and helicase domains of the dengue virus NS3 protein conferred by the linker region and its functional implications
Abstract
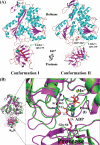
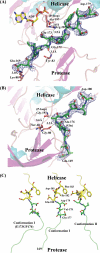
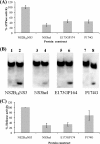
The dengue virus (DENV) NS3 protein is essential for viral polyprotein processing and RNA replication. It contains an N-terminal serine protease region (residues 1-168) joined to an RNA helicase (residues 180-618) by an 11-amino acid linker (169-179). The structure at 3.15 A of the soluble NS3 protein from DENV4 covalently attached to 18 residues of the NS2B cofactor region (NS2B(18)NS3) revealed an elongated molecule with the protease domain abutting subdomains I and II of the helicase (Luo, D., Xu, T., Hunke, C., Grüber, G., Vasudevan, S. G., and Lescar, J. (2008) J. Virol. 82, 173-183). Unexpectedly, using similar crystal growth conditions, we observed an alternative conformation where the protease domain has rotated by approximately 161 degrees with respect to the helicase domain. We report this new crystal structure bound to ADP-Mn(2+) refined to a resolution of 2.2 A. The biological significance for interdomain flexibility conferred by the linker region was probed by either inserting a Gly residue between Glu(173) and Pro(174) or replacing Pro(174) with a Gly residue. Both mutations resulted in significantly lower ATPase and helicase activities. We next increased flexibility in the linker by introducing a Pro(176) to Gly mutation in a DENV2 replicon system. A 70% reduction in luciferase reporter signal and a similar reduction in the level of viral RNA synthesis were observed. Our results indicate that the linker region has evolved to an optimum length to confer flexibility to the NS3 protein that is required both for polyprotein processing and RNA replication.
Figures

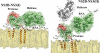
Similar articles
-
Dengue virus serotypic replacement of NS3 protease or helicase domain causes chimeric viral attenuation but can be recovered by a compensated mutation at helicase domain or NS2B, respectively.J Virol. 2023 Aug 31;97(8):e0085423. doi: 10.1128/jvi.00854-23. Epub 2023 Aug 9. J Virol. 2023. PMID: 37555662 Free PMC article.
-
Uncoupling of Protease trans-Cleavage and Helicase Activities in Pestivirus NS3.J Virol. 2017 Oct 13;91(21):e01094-17. doi: 10.1128/JVI.01094-17. Print 2017 Nov 1. J Virol. 2017. PMID: 28835495 Free PMC article.
-
Crystal structures of full length DENV4 NS2B-NS3 reveal the dynamic interaction between NS2B and NS3.Antiviral Res. 2020 Oct;182:104900. doi: 10.1016/j.antiviral.2020.104900. Epub 2020 Aug 5. Antiviral Res. 2020. PMID: 32763315
-
Structure and flexibility of non-structural proteins 3 and -5 of Dengue- and Zika viruses in solution.Prog Biophys Mol Biol. 2019 May;143:67-77. doi: 10.1016/j.pbiomolbio.2018.08.008. Epub 2018 Aug 29. Prog Biophys Mol Biol. 2019. PMID: 30171868 Review.
-
The flavivirus NS2B-NS3 protease-helicase as a target for antiviral drug development.Antiviral Res. 2015 Jun;118:148-58. doi: 10.1016/j.antiviral.2015.03.014. Epub 2015 Apr 2. Antiviral Res. 2015. PMID: 25842996 Review.
Cited by
-
In-silico screening for anti-Zika virus phytochemicals.J Mol Graph Model. 2016 Sep;69:78-91. doi: 10.1016/j.jmgm.2016.08.011. Epub 2016 Aug 28. J Mol Graph Model. 2016. PMID: 27588363 Free PMC article.
-
Role of Endoplasmic Reticulum-Associated Proteins in Flavivirus Replication and Assembly Complexes.Pathogens. 2019 Sep 12;8(3):148. doi: 10.3390/pathogens8030148. Pathogens. 2019. PMID: 31547236 Free PMC article. Review.
-
Potential Role of Flavivirus NS2B-NS3 Proteases in Viral Pathogenesis and Anti-flavivirus Drug Discovery Employing Animal Cells and Models: A Review.Viruses. 2021 Dec 28;14(1):44. doi: 10.3390/v14010044. Viruses. 2021. PMID: 35062249 Free PMC article. Review.
-
A novel dengue virus inhibitor, BP13944, discovered by high-throughput screening with dengue virus replicon cells selects for resistance in the viral NS2B/NS3 protease.Antimicrob Agents Chemother. 2014;58(1):110-9. doi: 10.1128/AAC.01281-13. Epub 2013 Oct 21. Antimicrob Agents Chemother. 2014. PMID: 24145533 Free PMC article.
-
Organization of the Flavivirus RNA replicase complex.Wiley Interdiscip Rev RNA. 2017 Nov;8(6):10.1002/wrna.1437. doi: 10.1002/wrna.1437. Epub 2017 Aug 16. Wiley Interdiscip Rev RNA. 2017. PMID: 28815931 Free PMC article. Review.
References
-
- Lindenbach B. D., Thiel H. J., Rice C. M. (2007) in Fields Virology (Knipe D. M., Howley P. M. eds) 5th Ed., pp. 1101–1152, Lippincott-Raven Publishers, Philadelphia
-
- Erbel P., Schiering N., D'Arcy A., Renatus M., Kroemer M., Lim S. P., Yin Z., Keller T. H., Vasudevan S. G., Hommel U. (2006) Nat. Struct. Mol. Biol. 13, 372–373 - PubMed
-
- Robin G., Chappell K., Stoermer M. J., Hu S. H., Young P. R., Fairlie D. P., Martin J. L. (2009) J. Mol. Biol. 385, 1568–1577 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

