Phylogeography of two moray eels indicates high dispersal throughout the indo-pacific
- PMID: 20375076
- PMCID: PMC2884193
- DOI: 10.1093/jhered/esq036
Phylogeography of two moray eels indicates high dispersal throughout the indo-pacific
Abstract
Reef fishes disperse primarily as oceanic "pelagic" larvae, and debate continues over the extent of this dispersal, with recent evidence for geographically restricted (closed) populations in some species. In contrast, moray eels have the longest pelagic larval stages among reef fishes, possibly providing opportunities to disperse over great distances. We test this prediction by measuring mitochondrial DNA (mtDNA) and nuclear DNA variation in 2 species of moray eels, Gymnothorax undulatus (N = 165) and G. flavimarginatus (N = 124), sampled at 14-15 locations across the Indo-Pacific. The mtDNA data comprise 632 bp of cytochrome b and 596 bp of cytochrome oxidase I. Nuclear markers include 2 recombination-activating loci (421 bp of RAG-1 and 754 bp of RAG-2). Analyses of molecular variance and Mantel tests indicate little or no genetic differentiation, and no isolation by distance, across 22 000 km of the Indo-Pacific. We estimate that mitochondrial genetic variation coalesces within the past about 2.3 million years (My) for G. flavimarginatus and within the past about 5.9 My for G. undulatus. Permutation tests of geographic distance on the mitochondrial haplotype networks indicate recent range expansions for some younger haplotypes (estimated within approximately 600 000 years) and episodic fragmentation of populations at times of low sea level. Our results support the predictions that the extended larval durations of moray eels enable ocean-wide genetic continuity of populations. This is the first phylogeographic survey of the moray eels, and morays are the first reef fishes known to be genetically homogeneous across the entire Indo-Pacific.
Figures

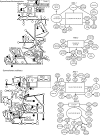
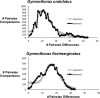
References
-
- Atema J, Kingsford MJ, Gerlack G. Larval reef fish could use odour for detection, retention and orientation to reefs. Mar Ecol Prog Ser. 2002;241:151–160.
-
- Barber PH, Erdmann MV, Palumbi SR. Comparative phylogeography of three codistributed stomatopods: origins and timing of regional lineage diversification in the coral triangle. Evolution. 2006;60:1825–1839. - PubMed
-
- Bay LK, Choat HJ, Van Herwerden L, Robertson DR. High genetic diversities and complex genetic structure in an Indo-Pacific tropical reef fish (Chlorurus sordidus): evidence of an unstable evolutionary past? Mar Biol. 2004;144:757–767.
-
- Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Proc B. 1995;57:289–300.
-
- Bermingham E, McCafferty SS, Martin AP. Fish biogeography and molecular clocks: perspectives from the Panamanian isthmus. In: Kocher TD, Stepien CA, editors. Molecular systematics of fishes. New York: Academic Press; 1997. pp. 113–128.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

