The role of the universally conserved A2450-C2063 base pair in the ribosomal peptidyl transferase center
- PMID: 20375101
- PMCID: PMC2919715
- DOI: 10.1093/nar/gkq213
The role of the universally conserved A2450-C2063 base pair in the ribosomal peptidyl transferase center
Abstract
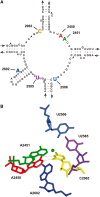
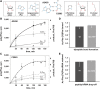
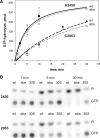
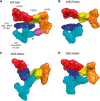
Despite the fact that all 23S rRNA nucleotides that build the ribosomal peptidyl transferase ribozyme are universally conserved, standard and atomic mutagenesis studies revealed the nucleobase identities being non-critical for catalysis. This indicates that these active site residues are highly conserved for functions distinct from catalysis. To gain insight into potential contributions, we have manipulated the nucleobases via an atomic mutagenesis approach and have utilized these chemically engineered ribosomes for in vitro translation reactions. We show that most of the active site nucleobases could be removed without significant effects on polypeptide production. Our data however highlight the functional importance of the universally conserved non-Watson-Crick base pair at position A2450-C2063. Modifications that disrupt this base pair markedly impair translation activities, while having little effects on peptide bond formation, tRNA drop-off and ribosome-dependent EF-G GTPase activity. Thus it seems that disruption of the A2450-C2063 pair inhibits a reaction following transpeptidation and EF-G action during the elongation cycle. Cumulatively our data are compatible with the hypothesis that the integrity of this A-C wobble base pair is essential for effective tRNA translocation through the peptidyl transferase center during protein synthesis.
Figures


Similar articles
-
Exploration of the conserved A+C wobble pair within the ribosomal peptidyl transferase center using affinity purified mutant ribosomes.Nucleic Acids Res. 2004 Jul 15;32(12):3760-70. doi: 10.1093/nar/gkh672. Print 2004. Nucleic Acids Res. 2004. PMID: 15256541 Free PMC article.
-
An intact ribose moiety at A2602 of 23S rRNA is key to trigger peptidyl-tRNA hydrolysis during translation termination.Nucleic Acids Res. 2007;35(15):5130-40. doi: 10.1093/nar/gkm539. Epub 2007 Jul 26. Nucleic Acids Res. 2007. PMID: 17660192 Free PMC article.
-
The A2453-C2499 wobble base pair in Escherichia coli 23S ribosomal RNA is responsible for pH sensitivity of the peptidyltransferase active site conformation.Nucleic Acids Res. 2004 Oct 12;32(18):5512-8. doi: 10.1093/nar/gkh888. Print 2004. Nucleic Acids Res. 2004. PMID: 15479786 Free PMC article.
-
Importance of tRNA interactions with 23S rRNA for peptide bond formation on the ribosome: studies with substrate analogs.Biol Chem. 2007 Jul;388(7):687-91. doi: 10.1515/BC.2007.077. Biol Chem. 2007. PMID: 17570820 Review.
-
Peptidyl transferase and beyond.Biochem Cell Biol. 1995 Nov-Dec;73(11-12):1041-7. doi: 10.1139/o95-111. Biochem Cell Biol. 1995. PMID: 8722019 Review.
Cited by
-
Mutational characterization and mapping of the 70S ribosome active site.Nucleic Acids Res. 2020 Mar 18;48(5):2777-2789. doi: 10.1093/nar/gkaa001. Nucleic Acids Res. 2020. PMID: 32009164 Free PMC article.
-
Oxidative stress damages rRNA inside the ribosome and differentially affects the catalytic center.Nucleic Acids Res. 2018 Feb 28;46(4):1945-1957. doi: 10.1093/nar/gkx1308. Nucleic Acids Res. 2018. PMID: 29309687 Free PMC article.
-
Critical 23S rRNA interactions for macrolide-dependent ribosome stalling on the ErmCL nascent peptide chain.Nucleic Acids Res. 2017 Jun 20;45(11):6717-6728. doi: 10.1093/nar/gkx195. Nucleic Acids Res. 2017. PMID: 28369621 Free PMC article.
-
Generation of chemically engineered ribosomes for atomic mutagenesis studies on protein biosynthesis.Nat Protoc. 2011 May;6(5):580-92. doi: 10.1038/nprot.2011.306. Epub 2011 Apr 7. Nat Protoc. 2011. PMID: 21527916
-
Unifying the Aminohexopyranose- and Peptidyl-Nucleoside Antibiotics: Implications for Antibiotic Design.Angew Chem Int Ed Engl. 2020 Jul 6;59(28):11330-11333. doi: 10.1002/anie.202003094. Epub 2020 May 11. Angew Chem Int Ed Engl. 2020. PMID: 32342623 Free PMC article.
References
-
- Fox GE, Naik AK. The evolutionary history of the ribosome. In: de Pouplana LR, editor. The Genetic Code and the Origin of Life. Texas: Landes Bioscience; 2004. pp. 92–105.
-
- Erlacher MD, Polacek N. Ribosomal catalysis: the evolution of mechanistic concepts for peptide bond formation and peptidyl-tRNA hydrolysis. RNA Biol. 2008;5:5–12. - PubMed
-
- Maden BE. Historical review: peptidyl transfer, the Monro era. Trends Biochem. Sci. 2003;28:619–624. - PubMed
-
- Polacek N, Mankin AS. The ribosomal peptidyl transferase center: structure, function, evolution, inhibition. Crit. Rev. Biochem. Mol. 2005;40:285–311. - PubMed
-
- Youngman EM, Brunelle JL, Kochaniak AB, Green R. The active site of the ribosome is composed of two layers of conserved nucleotides with distinct roles in peptide bond formation and peptide release. Cell. 2004;117:589–599. - PubMed

