Degradation of human alpha- and beta-defensins by culture supernatants of Porphyromonas gingivalis strain 381
- PMID: 20375570
- PMCID: PMC7312840
- DOI: 10.1159/000181015
Degradation of human alpha- and beta-defensins by culture supernatants of Porphyromonas gingivalis strain 381
Abstract
Porphyromonas gingivalis produces proteases capable of degrading cytokines, host heme proteins and some antimicrobial peptides. In this study, we show that P. gingivalis culture supernatants fully or partially degrade human neutrophil peptide alpha-defensins and human beta-defensins after 30 min. This observation suggests that proteases from P. gingivalis degrade defensins and this activity could abrogate defensin-related innate immune functions.
Copyright 2008 S. Karger AG, Basel.
Figures
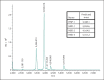
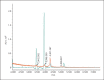
Comment in
-
Bacterial proteases disarming host defense.J Innate Immun. 2009;1(2):69. doi: 10.1159/000181143. Epub 2008 Dec 2. J Innate Immun. 2009. PMID: 20375567 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

