Promoter polymorphism -119C/G in MYG1 (C12orf10) gene is related to vitiligo susceptibility and Arg4Gln affects mitochondrial entrance of Myg1
- PMID: 20377893
- PMCID: PMC2856544
- DOI: 10.1186/1471-2350-11-56
Promoter polymorphism -119C/G in MYG1 (C12orf10) gene is related to vitiligo susceptibility and Arg4Gln affects mitochondrial entrance of Myg1
Abstract
Background: MYG1 (Melanocyte proliferating gene 1, also C12orf10 in human) is a ubiquitous nucleo-mitochondrial protein, involved in early developmental processes and in adult stress/illness conditions. We recently showed that MYG1 mRNA expression is elevated in the skin of vitiligo patients. Our aim was to examine nine known polymorphisms in the MYG1 gene, to investigate their functionality, and to study their association with vitiligo susceptibility.
Methods: Nine single nucleotide polymorphisms (SNPs) in the MYG1 locus were investigated by SNPlex assay and/or sequencing in vitiligo patients (n = 124) and controls (n = 325). MYG1 expression in skin biopsies was detected by quantitative-real time PCR (Q-RT-PCR) and polymorphisms were further analysed using luciferase and YFP reporters in the cell culture.
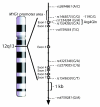
Results: Control subjects with -119G promoter allele (rs1465073) exhibited significantly higher MYG1 mRNA levels than controls with -119C allele (P = 0.01). Higher activity of -119G promoter was confirmed by luciferase assay. Single marker association analysis showed that the -119G allele was more frequent in vitiligo patients (47.1%) compared to controls (39.3%, P < 0.05, OR 1.37, 95%CI 1.02-1.85). Analysis based on the stage of progression of the vitiligo revealed that the increased frequency of -119G allele occurred prevalently in the group of patients with active vitiligo (n = 86) compared to the control group (48.2% versus 39.3%, P < 0.05; OR 1.44, 95%CI 1.02-2.03). Additionally, we showed that glutamine in the fourth position (in Arg4Gln polymorphism) completely eliminated mitochondrial entrance of YFP-tagged Myg1 protein in cell culture. The analysis of available EST, cDNA and genomic DNA sequences revealed that Myg1 4Gln allele is remarkably present in human populations but is never detected in homozygous state according to the HapMap database.
Conclusions: Our study demonstrated that both MYG1 promoter polymorphism -119C/G and Arg4Gln polymorphism in the mitochondrial signal of Myg1 have a functional impact on the regulation of the MYG1 gene and promoter polymorphism (-119C/G) is related with suspectibility for actively progressing vitiligo.
Figures
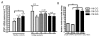


Similar articles
-
Correlation of increased MYG1 expression and its promoter polymorphism with disease progression and higher susceptibility in vitiligo patients.J Dermatol Sci. 2013 Sep;71(3):195-202. doi: 10.1016/j.jdermsci.2013.04.026. Epub 2013 May 1. J Dermatol Sci. 2013. PMID: 23706493
-
Association of FOXP3 and GAGE10 promoter polymorphisms and decreased FOXP3 expression in regulatory T cells with susceptibility to generalized vitiligo in Gujarat population.Gene. 2021 Feb 5;768:145295. doi: 10.1016/j.gene.2020.145295. Epub 2020 Nov 9. Gene. 2021. PMID: 33181260
-
Myg1 exonuclease couples the nuclear and mitochondrial translational programs through RNA processing.Nucleic Acids Res. 2019 Jun 20;47(11):5852-5866. doi: 10.1093/nar/gkz371. Nucleic Acids Res. 2019. PMID: 31081026 Free PMC article.
-
Association of ACE gene I/D polymorphism with vitiligo: a meta-analysis.Arch Dermatol Res. 2013 Jul;305(5):365-70. doi: 10.1007/s00403-013-1315-z. Epub 2013 Jan 17. Arch Dermatol Res. 2013. PMID: 23325447 Review.
-
Association of the 389 C/T polymorphism of the catalase gene with susceptibility to vitiligo: a meta-analysis.Clin Exp Dermatol. 2014 Jun;39(4):454-60. doi: 10.1111/ced.12340. Clin Exp Dermatol. 2014. PMID: 24825136 Review.
Cited by
-
Recent progress in the genetics of generalized vitiligo.J Genet Genomics. 2011 Jul 20;38(7):271-8. doi: 10.1016/j.jgg.2011.05.005. Epub 2011 Jun 12. J Genet Genomics. 2011. PMID: 21777851 Free PMC article. Review.
-
Modern vitiligo genetics sheds new light on an ancient disease.J Dermatol. 2013 May;40(5):310-8. doi: 10.1111/1346-8138.12147. J Dermatol. 2013. PMID: 23668538 Free PMC article. Review.
-
MYG1 promotes proliferation and inhibits autophagy in lung adenocarcinoma cells via the AMPK/mTOR complex 1 signaling pathway.Oncol Lett. 2021 Apr;21(4):334. doi: 10.3892/ol.2021.12595. Epub 2021 Feb 25. Oncol Lett. 2021. PMID: 33692866 Free PMC article.
-
Shared genetic relationships underlying generalized vitiligo and autoimmune thyroid disease.Thyroid. 2010 Jul;20(7):745-54. doi: 10.1089/thy.2010.1643. Thyroid. 2010. PMID: 20578892 Free PMC article. Review.
-
Update on the genetics characterization of vitiligo.Int J Health Sci (Qassim). 2011 Jul;5(2):167-79. Int J Health Sci (Qassim). 2011. PMID: 23267294 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

