Molecular targets of epigenetic regulation and effectors of environmental influences
- PMID: 20381512
- PMCID: PMC2911443
- DOI: 10.1016/j.taap.2010.03.022
Molecular targets of epigenetic regulation and effectors of environmental influences
Abstract
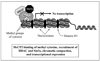
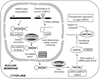
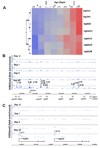
The true understanding of what we currently define as epigenetics evolved over time as our knowledge on DNA methylation and chromatin modifications and their effects on gene expression increased. The current explosion of research on epigenetics and the increasing documentation of the effects of various environmental factors on DNA methylation, chromatin modification, as well as on the expression of small non-coding RNAs (ncRNAs) have expanded the scope of research on the etiology of various diseases including cancer. The current review briefly discusses the molecular mechanisms of epigenetic regulation and expands the discussion with examples on the role of environment, such as the immediate environment during development, in inducing epigenetic changes and modulating gene expression.
Published by Elsevier Inc.
Conflict of interest statement
Figures



Similar articles
-
Epigenetic mechanisms in metal toxicity.Toxicol Mech Methods. 2011 May;21(4):343-52. doi: 10.3109/15376516.2011.557878. Toxicol Mech Methods. 2011. PMID: 21495872 Review.
-
Epigenetics and ocular diseases: from basic biology to clinical study.J Cell Physiol. 2014 Jul;229(7):825-33. doi: 10.1002/jcp.24522. J Cell Physiol. 2014. PMID: 24318407 Review.
-
Non-coding RNAs as direct and indirect modulators of epigenetic regulation.Epigenetics. 2014 Jan;9(1):3-12. doi: 10.4161/epi.27473. Epigenetics. 2014. PMID: 24739571 Free PMC article. Review.
-
Transcriptional gene silencing through epigenetic changes mediated by non-coding RNAs.Curr Opin Mol Ther. 2010 Apr;12(2):214-22. Curr Opin Mol Ther. 2010. PMID: 20373265 Free PMC article. Review.
-
Role of Non-Coding RNAs in the Transgenerational Epigenetic Transmission of the Effects of Reprotoxicants.Int J Mol Sci. 2016 Mar 25;17(4):452. doi: 10.3390/ijms17040452. Int J Mol Sci. 2016. PMID: 27023531 Free PMC article. Review.
Cited by
-
Maternal smoking as a model for environmental epigenetic changes affecting birthweight and fetal programming.Mol Hum Reprod. 2013 Jan;19(1):1-6. doi: 10.1093/molehr/gas050. Epub 2012 Nov 8. Mol Hum Reprod. 2013. PMID: 23139402 Free PMC article. Review.
-
The role of miRNAs 34a, 146a, 320a and 542 in the synergistic anticancer effects of methyl 2-(5-fluoro-2-hydroxyphenyl)-1H- benzo[d]imidazole-5-carboxylate (MBIC) with doxorubicin in breast cancer cells.PeerJ. 2018 Sep 17;6:e5577. doi: 10.7717/peerj.5577. eCollection 2018. PeerJ. 2018. PMID: 30245930 Free PMC article.
-
Effect of valproic acid upon skeletal muscle subjected to prolonged tourniquet application.Trauma Surg Acute Care Open. 2023 Jul 19;8(1):e001074. doi: 10.1136/tsaco-2022-001074. eCollection 2023. Trauma Surg Acute Care Open. 2023. PMID: 37484837 Free PMC article.
-
Breast cancer resistance protein BCRP/ABCG2 regulatory microRNAs (hsa-miR-328, -519c and -520h) and their differential expression in stem-like ABCG2+ cancer cells.Biochem Pharmacol. 2011 Mar 15;81(6):783-92. doi: 10.1016/j.bcp.2010.12.018. Epub 2011 Jan 8. Biochem Pharmacol. 2011. PMID: 21219875 Free PMC article.
-
Impact of Epigenetic Dietary Components on Cancer through Histone Modifications.Curr Med Chem. 2015;22(17):2051-64. doi: 10.2174/0929867322666150420102641. Curr Med Chem. 2015. PMID: 25891109 Free PMC article. Review.
References
-
- Alnouti Y, Patrick JS, Klaassen CD. Tissue distribution and ontogeny of organic cation transporter in mice. Drug. Metab. Disp. 2006;34:477–482. - PubMed
-
- Berger S. The complex language of chromatin regulation during transcription. Nature. 2007;447:407–412. - PubMed
-
- Bhaumik SR, Smith E, Shilatifard A. Covalent modifications of histones during development and disease pathogenesis. Nat. Struct. Mol. Biol. 2007;14:1008–1016. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

