An expanding universe of circadian networks in higher plants
- PMID: 20382065
- PMCID: PMC2866796
- DOI: 10.1016/j.tplants.2010.03.003
An expanding universe of circadian networks in higher plants
Abstract
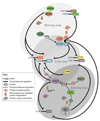
Extensive circadian clock networks regulate almost every biological process in plants. Clock-controlled physiological responses are coupled with daily oscillations in environmental conditions resulting in enhanced fitness and growth vigor. Identification of core clock components and their associated molecular interactions has established the basic network architecture of plant clocks, which consists of multiple interlocked feedback loops. A hierarchical structure of transcriptional feedback overlaid with regulated protein turnover sets the pace of the clock and ultimately drives all clock-controlled processes. Although originally described as linear entities, increasing evidence suggests that many signaling pathways can act as both inputs and outputs within the overall network. Future studies will determine the molecular mechanisms involved in these complex regulatory loops.
2010 Elsevier Ltd. All rights reserved.
Figures

References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

