Biosynthesis of wyosine derivatives in tRNA: an ancient and highly diverse pathway in Archaea
- PMID: 20382657
- PMCID: PMC4481705
- DOI: 10.1093/molbev/msq096
Biosynthesis of wyosine derivatives in tRNA: an ancient and highly diverse pathway in Archaea
Abstract
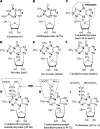
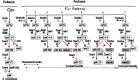
Wyosine (imG) and its derivatives such as wybutosine (yW) are found at position 37 of phenylalanine-specific transfer RNA (tRNA(Phe)), 3' adjacent to the anticodon in Eucarya and Archaea. In Saccharomyces cerevisiae, formation of yW requires five enzymes acting in a strictly sequential order: Trm5, Tyw1, Tyw2, Tyw3, and Tyw4. Archaea contain wyosine derivatives, but their diversity is greater than in eukaryotes and the corresponding biosynthesis pathways still unknown. To identify these pathways, we analyzed the phylogenetic distribution of homologues of the yeast wybutosine biosynthesis proteins in 62 archaeal genomes and proposed a scenario for the origin and evolution of wyosine derivatives biosynthesis in Archaea that was partly experimentally validated. The key observations were 1) that four of the five wybutosine biosynthetic enzymes are ancient and may have been present in the last common ancestor of Archaea and Eucarya, 2) that the variations in the distribution pattern of biosynthesis enzymes reflect the diversity of the wyosine derivatives found in different Archaea. We also identified 7-aminocarboxypropyl-demethylwyosine (yW-86) and its N4-methyl derivative (yW-72) as final products in tRNAs of several Archaea when these were previously thought to be only intermediates of the eukaryotic pathway. We confirmed that isowyosine (imG2) and 7-methylwyosine (mimG) are two archaeal-specific guanosine-37 derivatives found in tRNA of both Euryarchaeota and Crenarchaeota. Finally, we proposed that the duplication of the trm5 gene in some Archaea led to a change in function from N1 methylation of guanosine to C7 methylation of 4-demethylwyosine (imG-14).
Figures




References
-
- Altschul SF, Koonin EV. Iterated profile searches with PSI-BLAST—a tool for discovery in protein databases. Trends Biochem Sci. 1998;23:444. - PubMed
-
- Björk GR, Hagervall TG. Transfer RNA modification. In: Böck A, Curtis R, Kaper JB, Neidhardt FC, Nyström T, Squires CL, editors. Escherichia coliand Salmonella. Cellular and molecular biology. Washington (DC): ASM Press; 2005. [cited 2010 April 29]. Available from: http://www.ecosal.org.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

