Shape based kinetic outlier detection in real-time PCR
- PMID: 20385019
- PMCID: PMC2873533
- DOI: 10.1186/1471-2105-11-186
Shape based kinetic outlier detection in real-time PCR
Abstract
Background: Real-time PCR has recently become the technique of choice for absolute and relative nucleic acid quantification. The gold standard quantification method in real-time PCR assumes that the compared samples have similar PCR efficiency. However, many factors present in biological samples affect PCR kinetic, confounding quantification analysis. In this work we propose a new strategy to detect outlier samples, called SOD.
Results: Richards function was fitted on fluorescence readings to parameterize the amplification curves. There was not a significant correlation between calculated amplification parameters (plateau, slope and y-coordinate of the inflection point) and the Log of input DNA demonstrating that this approach can be used to achieve a "fingerprint" for each amplification curve. To identify the outlier runs, the calculated parameters of each unknown sample were compared to those of the standard samples. When a significant underestimation of starting DNA molecules was found, due to the presence of biological inhibitors such as tannic acid, IgG or quercitin, SOD efficiently marked these amplification profiles as outliers. SOD was subsequently compared with KOD, the current approach based on PCR efficiency estimation. The data obtained showed that SOD was more sensitive than KOD, whereas SOD and KOD were equally specific.
Conclusion: Our results demonstrated, for the first time, that outlier detection can be based on amplification shape instead of PCR efficiency. SOD represents an improvement in real-time PCR analysis because it decreases the variance of data thus increasing the reliability of quantification.
Figures



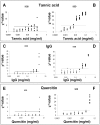
 ; where
; where  is the mean of parameter calculated for each inhibitor concentration;
is the mean of parameter calculated for each inhibitor concentration;  represents the mean of parameter value from standard curve samples (C). Right lower panel: asymmetry values versus increasing inhibitor concentration. Asymmetry was computed as the following ratio:
represents the mean of parameter value from standard curve samples (C). Right lower panel: asymmetry values versus increasing inhibitor concentration. Asymmetry was computed as the following ratio:  (D).
(D).

Similar articles
-
Kinetic Outlier Detection (KOD) in real-time PCR.Nucleic Acids Res. 2003 Sep 1;31(17):e105. doi: 10.1093/nar/gng106. Nucleic Acids Res. 2003. PMID: 12930979 Free PMC article.
-
Validation of kinetics similarity in qPCR.Nucleic Acids Res. 2012 Feb;40(4):1395-406. doi: 10.1093/nar/gkr778. Epub 2011 Oct 19. Nucleic Acids Res. 2012. PMID: 22013160 Free PMC article. Review.
-
A kinetic-based sigmoidal model for the polymerase chain reaction and its application to high-capacity absolute quantitative real-time PCR.BMC Biotechnol. 2008 May 8;8:47. doi: 10.1186/1472-6750-8-47. BMC Biotechnol. 2008. PMID: 18466619 Free PMC article.
-
A new real-time PCR method to overcome significant quantitative inaccuracy due to slight amplification inhibition.BMC Bioinformatics. 2008 Jul 30;9:326. doi: 10.1186/1471-2105-9-326. BMC Bioinformatics. 2008. PMID: 18667053 Free PMC article.
-
[Quantitative PCR in the diagnosis of Leishmania].Parassitologia. 2004 Jun;46(1-2):163-7. Parassitologia. 2004. PMID: 15305709 Review. Italian.
Cited by
-
Specific serum and CSF microRNA profiles distinguish sporadic behavioural variant of frontotemporal dementia compared with Alzheimer patients and cognitively healthy controls.PLoS One. 2018 May 10;13(5):e0197329. doi: 10.1371/journal.pone.0197329. eCollection 2018. PLoS One. 2018. PMID: 29746584 Free PMC article.
-
Enhanced analysis of real-time PCR data by using a variable efficiency model: FPK-PCR.Nucleic Acids Res. 2012 Jan;40(2):e10. doi: 10.1093/nar/gkr775. Epub 2011 Nov 18. Nucleic Acids Res. 2012. PMID: 22102586 Free PMC article.
-
Effect of Different Exercise Intensities on the Myotendinous Junction Plasticity.PLoS One. 2016 Jun 23;11(6):e0158059. doi: 10.1371/journal.pone.0158059. eCollection 2016. PLoS One. 2016. PMID: 27337061 Free PMC article.
-
Framework for DNA Quantification and Outlier Detection Using Multidimensional Standard Curves.Anal Chem. 2019 Jun 4;91(11):7426-7434. doi: 10.1021/acs.analchem.9b01466. Epub 2019 May 14. Anal Chem. 2019. PMID: 31056898 Free PMC article.
-
Empirical evaluation of humpback whale telomere length estimates; quality control and factors causing variability in the singleplex and multiplex qPCR methods.BMC Genet. 2012 Sep 6;13:77. doi: 10.1186/1471-2156-13-77. BMC Genet. 2012. PMID: 22954451 Free PMC article.
References
-
- VanGuilder HD, Vrana KE, Freeman WM. Twenty-five years of quantitative PCR for gene expression analysis. Bio Techniques. 2008;44(5):619–626. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

