Continued primer synthesis at stalled replication forks contributes to checkpoint activation
- PMID: 20385778
- PMCID: PMC2856894
- DOI: 10.1083/jcb.200909105
Continued primer synthesis at stalled replication forks contributes to checkpoint activation
Abstract
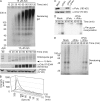
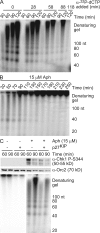
Stalled replication forks activate and are stabilized by the ATR (ataxia-telangiectasia mutated and Rad3 related)-mediated checkpoint, but ultimately, they must also recover from the arrest. Although primed single-stranded DNA (ssDNA) is sufficient for checkpoint activation, it is still unknown how this signal is generated at a stalled replication fork. Furthermore, it is not clear how recovery and fork restart occur in higher eukaryotes. Using Xenopus laevis egg extracts, we show that DNA replication continues at a stalled fork through the synthesis and elongation of new primers independent of the checkpoint. This synthesis is dependent on the activity of proliferating cell nuclear antigen, Pol-delta, and Pol-epsilon, and it contributes to the phosphorylation of Chk1. We also used defined DNA structures to show that for a fixed amount of ssDNA, increasing the number of primer-template junctions strongly enhances Chk1 phosphorylation. These results suggest that new primers are synthesized at stalled replication forks by the leading and lagging strand polymerases and that accumulation of these primers may contribute to checkpoint activation.
Figures





Similar articles
-
Functional uncoupling of MCM helicase and DNA polymerase activities activates the ATR-dependent checkpoint.Genes Dev. 2005 May 1;19(9):1040-52. doi: 10.1101/gad.1301205. Epub 2005 Apr 15. Genes Dev. 2005. PMID: 15833913 Free PMC article.
-
TopBP1 and DNA polymerase-alpha directly recruit the 9-1-1 complex to stalled DNA replication forks.J Cell Biol. 2009 Mar 23;184(6):793-804. doi: 10.1083/jcb.200810185. Epub 2009 Mar 16. J Cell Biol. 2009. PMID: 19289795 Free PMC article.
-
Phosphorylation of replication protein A by S-phase checkpoint kinases.DNA Repair (Amst). 2006 Mar 7;5(3):369-80. doi: 10.1016/j.dnarep.2005.11.007. Epub 2006 Jan 18. DNA Repair (Amst). 2006. PMID: 16412704
-
TopBP1 and DNA polymerase alpha-mediated recruitment of the 9-1-1 complex to stalled replication forks: implications for a replication restart-based mechanism for ATR checkpoint activation.Cell Cycle. 2009 Sep 15;8(18):2877-84. doi: 10.4161/cc.8.18.9485. Epub 2009 Sep 9. Cell Cycle. 2009. PMID: 19652550 Review.
-
ATR/Mec1: coordinating fork stability and repair.Curr Opin Cell Biol. 2009 Apr;21(2):237-44. doi: 10.1016/j.ceb.2009.01.017. Epub 2009 Feb 21. Curr Opin Cell Biol. 2009. PMID: 19230642 Review.
Cited by
-
Managing Single-Stranded DNA during Replication Stress in Fission Yeast.Biomolecules. 2015 Sep 18;5(3):2123-39. doi: 10.3390/biom5032123. Biomolecules. 2015. PMID: 26393661 Free PMC article. Review.
-
Maintenance of Genome Integrity: How Mammalian Cells Orchestrate Genome Duplication by Coordinating Replicative and Specialized DNA Polymerases.Genes (Basel). 2017 Jan 6;8(1):19. doi: 10.3390/genes8010019. Genes (Basel). 2017. PMID: 28067843 Free PMC article. Review.
-
DNA structure-specific priming of ATR activation by DNA-PKcs.J Cell Biol. 2013 Aug 5;202(3):421-9. doi: 10.1083/jcb.201304139. Epub 2013 Jul 29. J Cell Biol. 2013. PMID: 23897887 Free PMC article.
-
Mechanisms of loading and release of the 9-1-1 checkpoint clamp.Nat Struct Mol Biol. 2022 Apr;29(4):369-375. doi: 10.1038/s41594-022-00741-7. Epub 2022 Mar 21. Nat Struct Mol Biol. 2022. PMID: 35314831 Free PMC article.
-
Hsk1 kinase and Cdc45 regulate replication stress-induced checkpoint responses in fission yeast.Cell Cycle. 2010 Dec 1;9(23):4627-37. doi: 10.4161/cc.9.23.13937. Epub 2010 Dec 1. Cell Cycle. 2010. PMID: 21099360 Free PMC article.
References
-
- Bermudez V.P., Lindsey-Boltz L.A., Cesare A.J., Maniwa Y., Griffith J.D., Hurwitz J., Sancar A. 2003. Loading of the human 9-1-1 checkpoint complex onto DNA by the checkpoint clamp loader hRad17-replication factor C complex in vitro. Proc. Natl. Acad. Sci. USA. 100:1633–1638 10.1073/pnas.0437927100 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

