RASCAL is a new human cytomegalovirus-encoded protein that localizes to the nuclear lamina and in cytoplasmic vesicles at late times postinfection
- PMID: 20392852
- PMCID: PMC2903262
- DOI: 10.1128/JVI.02462-09
RASCAL is a new human cytomegalovirus-encoded protein that localizes to the nuclear lamina and in cytoplasmic vesicles at late times postinfection
Abstract
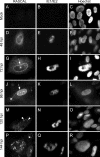
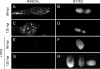
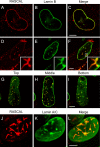
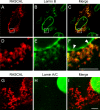
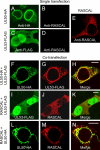
The products of numerous open reading frames (ORFs) present in the genome of human cytomegalovirus (CMV) have not been characterized. Here, we describe the identification of a new CMV protein localizing to the nuclear envelope and in cytoplasmic vesicles at late times postinfection. Based on this distinctive localization pattern, we called this new protein nuclear rim-associated cytomegaloviral protein, or RASCAL. Two RASCAL isoforms exist, a short version of 97 amino acids encoded by the majority of CMV strains and a longer version of 176 amino acids encoded by the Towne, Toledo, HAN20, and HAN38 strains. Both isoforms colocalize with lamin B in deep intranuclear invaginations of the inner nuclear membrane (INM) and in novel cytoplasmic vesicular structures possibly derived from the nuclear envelope. INM infoldings have been previously described as sites of nucleocapsid egress, which is mediated by the localized disruption of the nuclear lamina, promoted by the activities of viral and cellular kinases recruited by the lamina-associated proteins UL50 and UL53. RASCAL accumulation at the nuclear membrane required the presence of UL50 but not of UL53. RASCAL and UL50 also appeared to specifically interact, suggesting that RASCAL is a new component of the nuclear egress complex (NEC) and possibly involved in mediating nucleocapsid egress from the nucleus. Finally, the presence of RASCAL within cytoplasmic vesicles raises the intriguing possibility that this protein might participate in additional steps of virion maturation occurring after capsid release from the nucleus.
Figures


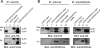
Similar articles
-
The absence of p53 during Human Cytomegalovirus infection leads to decreased UL53 expression, disrupting UL50 localization to the inner nuclear membrane, and thereby inhibiting capsid nuclear egress.Virology. 2016 Oct;497:262-278. doi: 10.1016/j.virol.2016.07.020. Epub 2016 Aug 4. Virology. 2016. PMID: 27498409 Free PMC article.
-
Human cytomegalovirus UL50 and UL53 recruit viral protein kinase UL97, not protein kinase C, for disruption of nuclear lamina and nuclear egress in infected cells.J Virol. 2014 Jan;88(1):249-62. doi: 10.1128/JVI.02358-13. Epub 2013 Oct 23. J Virol. 2014. PMID: 24155370 Free PMC article.
-
Human cytomegalovirus UL97 phosphorylates the viral nuclear egress complex.J Virol. 2015 Jan;89(1):523-34. doi: 10.1128/JVI.02426-14. Epub 2014 Oct 22. J Virol. 2015. PMID: 25339763 Free PMC article.
-
Breach of the nuclear lamina during assembly of herpes simplex viruses.Nucleus. 2011 Jul-Aug;2(4):271-6. doi: 10.4161/nucl.2.4.16334. Epub 2011 Jul 1. Nucleus. 2011. PMID: 21941110 Free PMC article. Review.
-
Nuclear Egress of Herpesviruses: The Prototypic Vesicular Nucleocytoplasmic Transport.Adv Virus Res. 2016;94:81-140. doi: 10.1016/bs.aivir.2015.10.002. Epub 2016 Jan 29. Adv Virus Res. 2016. PMID: 26997591 Review.
Cited by
-
Viral and host control of cytomegalovirus maturation.Trends Microbiol. 2012 Aug;20(8):392-401. doi: 10.1016/j.tim.2012.04.008. Epub 2012 May 23. Trends Microbiol. 2012. PMID: 22633075 Free PMC article. Review.
-
The absence of p53 during Human Cytomegalovirus infection leads to decreased UL53 expression, disrupting UL50 localization to the inner nuclear membrane, and thereby inhibiting capsid nuclear egress.Virology. 2016 Oct;497:262-278. doi: 10.1016/j.virol.2016.07.020. Epub 2016 Aug 4. Virology. 2016. PMID: 27498409 Free PMC article.
-
Deletion of the human cytomegalovirus US17 gene increases the ratio of genomes per infectious unit and alters regulation of immune and endoplasmic reticulum stress response genes at early and late times after infection.J Virol. 2014 Feb;88(4):2168-82. doi: 10.1128/JVI.02704-13. Epub 2013 Dec 11. J Virol. 2014. PMID: 24335296 Free PMC article.
-
WDR5 Facilitates Human Cytomegalovirus Replication by Promoting Capsid Nuclear Egress.J Virol. 2018 Apr 13;92(9):e00207-18. doi: 10.1128/JVI.00207-18. Print 2018 May 1. J Virol. 2018. PMID: 29437978 Free PMC article.
-
Orchestration of protein acetylation as a toggle for cellular defense and virus replication.Nat Commun. 2018 Nov 23;9(1):4967. doi: 10.1038/s41467-018-07179-w. Nat Commun. 2018. PMID: 30470744 Free PMC article.
References
-
- Bendtsen, J. D., H. Nielsen, G. von Heijne, and S. Brunak. 2004. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 340:783-795. - PubMed
-
- Blom, N., T. Sicheritz-Ponten, R. Gupta, S. Gammeltoft, and S. Brunak. 2004. Prediction of post-translational glycosylation and phosphorylation of proteins from the amino acid sequence. Proteomics 4:1633-1649. - PubMed
-
- Britt, W. 2008. Manifestations of human cytomegalovirus infection: proposed mechanisms of acute and chronic disease. Curr. Top. Microbiol. Immunol. 325:417-470. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

