Biases in Illumina transcriptome sequencing caused by random hexamer priming
- PMID: 20395217
- PMCID: PMC2896536
- DOI: 10.1093/nar/gkq224
Biases in Illumina transcriptome sequencing caused by random hexamer priming
Abstract
Generation of cDNA using random hexamer priming induces biases in the nucleotide composition at the beginning of transcriptome sequencing reads from the Illumina Genome Analyzer. The bias is independent of organism and laboratory and impacts the uniformity of the reads along the transcriptome. We provide a read count reweighting scheme, based on the nucleotide frequencies of the reads, that mitigates the impact of the bias.
Figures


 . (b) As in (a), but the hexamers correspond to positions 25–30 of the aligned reads, with a correlation of
. (b) As in (a), but the hexamers correspond to positions 25–30 of the aligned reads, with a correlation of  .
.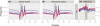
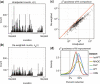
 goodness-of-fit statistics based on unadjusted and reweighted counts for 552 highly expressed regions of constant expression. (d) Smoothed histograms of the reduction in
goodness-of-fit statistics based on unadjusted and reweighted counts for 552 highly expressed regions of constant expression. (d) Smoothed histograms of the reduction in  goodness-of-fit statistics when using the re-weighting scheme, evaluated in five different experiments. Values greater than zero indicate that the re-weighting scheme improves the uniformity of the read distribution.
goodness-of-fit statistics when using the re-weighting scheme, evaluated in five different experiments. Values greater than zero indicate that the re-weighting scheme improves the uniformity of the read distribution.References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

