The spatial dynamics of tissue-specific promoters during C. elegans development
- PMID: 20395364
- PMCID: PMC2854392
- DOI: 10.1101/gad.559610
The spatial dynamics of tissue-specific promoters during C. elegans development
Abstract
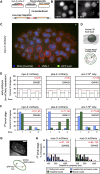
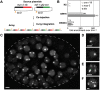
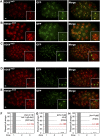
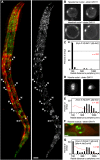
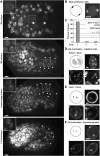
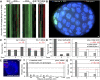
To understand whether the spatial organization of the genome reflects the cell's differentiated state, we examined whether genes assume specific subnuclear positions during Caenorhabditis elegans development. Monitoring the radial position of developmentally controlled promoters in embryos and larval tissues, we found that small integrated arrays bearing three different tissue-specific promoters have no preferential position in nuclei of undifferentiated embryos. However, in differentiated cells, they shifted stably toward the nuclear lumen when activated, or to the nuclear envelope when silent. In contrast, large integrated arrays bearing the same promoters became heterochromatic and nuclear envelope-bound in embryos. Tissue-specific activation of promoters in these large arrays in larvae overrode the perinuclear anchorage. For transgenes that carry both active and inactive promoters, the inward shift of the active promoter was dominant. Finally, induction of master regulator HLH-1 prematurely induced internalization of a muscle-specific promoter array in embryos. Fluorescence in situ hybridization confirmed analogous results for the endogenous endoderm-determining gene pha-4. We propose that, in differentiated cells, subnuclear organization arises from the selective positioning of active and inactive developmentally regulated promoters. We characterize two forces that lead to tissue-specific subnuclear organization of the worm genome: large repeat-induced heterochromatin, which associates with the nuclear envelope like repressed genes in differentiated cells, and tissue-specific promoters that shift inward in a dominant fashion over silent promoters, when they are activated.
Figures


References
-
- Akhtar A, Gasser SM 2007. The nuclear envelope and transcriptional control. Nat Rev Genet 8: 507–517 - PubMed
-
- Andersen EC, Horvitz HR 2007. Two C. elegans histone methyltransferases repress lin-3 EGF transcription to inhibit vulval development. Development 134: 2991–2999 - PubMed
-
- Azzaria M, Goszczynski B, Chung MA, Kalb JM, McGhee JD 1996. A fork head/HNF-3 homolog expressed in the pharynx and intestine of the Caenorhabditis elegans embryo. Dev Biol 178: 289–303 - PubMed
-
- Baugh LR, Hill AA, Claggett JM, Hill-Harfe K, Wen JC, Slonim DK, Brown EL, Hunter CP 2005. The homeodomain protein PAL-1 specifies a lineage-specific regulatory network in the C. elegans embryo. Development 132: 1843–1854 - PubMed
-
- Belmont AS 2001. Visualizing chromosome dynamics with GFP. Trends Cell Biol 11: 250–257 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
