Housekeeping gene stability influences the quantification of osteogenic markers during stem cell differentiation to the osteogenic lineage
- PMID: 20396946
- PMCID: PMC2873986
- DOI: 10.1007/s10616-010-9265-1
Housekeeping gene stability influences the quantification of osteogenic markers during stem cell differentiation to the osteogenic lineage
Abstract
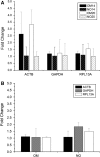
Real-time reverse transcription PCR (RT-qPCR) relies on a housekeeping or normalizer gene whose expression remains constant throughout the experiment. RT-qPCR is commonly used for characterization of human bone marrow mesenchymal stem cells (hBMSCs). However, to the best of our knowledge, there are no studies validating the expression stability of the genes used as normalizers during hBMSCs differentiation. This work aimed to study the stability of the housekeeping genes beta-actin, glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and ribosomal protein L13A (RPL13A) during the osteogenic differentiation of hBMSCs. Their stability was evaluated via RT-qPCR in 14 and 20 day differentiation assays to the osteogenic lineage. Different normalization strategies were evaluated to quantify the osteogenic markers collagen type I, bone sialoprotein and osteonectin. Cell differentiation was confirmed via alizarin red staining. The results demonstrated up-regulation of beta-actin with maximum fold changes (MFC) of 4.38. GAPDH and RPL13A were not regulated by osteogenic media after 14 days and presented average fold changes lower than 2 in 20 day cultures. RPL13A (MFC < 2) had a greater stability when normalizing as a function of culture time compared with GAPDH (MFC </= 2.2), which resulted in expression patterns of the osteogenic markers more consistent with the observed differentiation process. The results suggest that beta-actin regulation could be associated with the morphological changes characteristic of hBMSCs osteogenic differentiation, and provide evidence for the superior performance of RPL13A as a normalizer gene in osteogenic differentiation studies of hBMSCs. This work highlights the importance of validating the normalizer genes used for stem cells characterization via RT-qPCR.
Figures




Similar articles
-
EF1α is a suitable housekeeping gene for RT-qPCR analysis during osteogenic differentiation of mouse bone marrow-derived mesenchymal stem cells.Acta Biochim Pol. 2013;60(3):381-6. Acta Biochim Pol. 2013. PMID: 24051438
-
EF1alpha and RPL13a represent normalization genes suitable for RT-qPCR analysis of bone marrow derived mesenchymal stem cells.BMC Mol Biol. 2010 Aug 17;11:61. doi: 10.1186/1471-2199-11-61. BMC Mol Biol. 2010. PMID: 20716364 Free PMC article.
-
Effects of miR-103 by negatively regulating SATB2 on proliferation and osteogenic differentiation of human bone marrow mesenchymal stem cells.PLoS One. 2020 May 7;15(5):e0232695. doi: 10.1371/journal.pone.0232695. eCollection 2020. PLoS One. 2020. PMID: 32379794 Free PMC article.
-
Psoralen accelerates osteogenic differentiation of human bone marrow mesenchymal stem cells by activating the TGF-β/Smad3 pathway.Exp Ther Med. 2021 Sep;22(3):940. doi: 10.3892/etm.2021.10372. Epub 2021 Jul 1. Exp Ther Med. 2021. PMID: 34306204 Free PMC article.
-
MiR-138-5p knockdown promotes osteogenic differentiation through FOXC1 up-regulation in human bone mesenchymal stem cells.Biochem Cell Biol. 2021 Jun;99(3):296-303. doi: 10.1139/bcb-2020-0163. Epub 2020 Oct 15. Biochem Cell Biol. 2021. PMID: 33058690
Cited by
-
Gapdh gene expression is modulated by inflammatory arthritis and is not suitable for qPCR normalization.Inflammation. 2014 Aug;37(4):1059-69. doi: 10.1007/s10753-014-9829-x. Inflammation. 2014. PMID: 24493325
-
RPLP0/TBP are the most stable reference genes for human dental pulp stem cells under osteogenic differentiation.World J Stem Cells. 2024 Jun 26;16(6):656-669. doi: 10.4252/wjsc.v16.i6.656. World J Stem Cells. 2024. PMID: 38948092 Free PMC article.
-
RPL13A and EEF1A1 Are Suitable Reference Genes for qPCR during Adipocyte Differentiation of Vascular Stromal Cells from Patients with Different BMI and HOMA-IR.PLoS One. 2016 Jun 15;11(6):e0157002. doi: 10.1371/journal.pone.0157002. eCollection 2016. PLoS One. 2016. PMID: 27304673 Free PMC article.
-
Reference loci for RT-qPCR analysis of differentiating human embryonic stem cells.BMC Mol Biol. 2013 Sep 12;14:21. doi: 10.1186/1471-2199-14-21. BMC Mol Biol. 2013. PMID: 24028740 Free PMC article.
-
Markers Are Shared Between Adipogenic and Osteogenic Differentiated Mesenchymal Stem Cells.J Dev Biol Tissue Eng. 2013 May 1;5(2):18-25. doi: 10.5897/JDBTE2013.0065. J Dev Biol Tissue Eng. 2013. PMID: 24013643 Free PMC article.
References
-
- Abdallah BM, Haack-Sorensen M, Burns JS, Elsnab B, Jakob F, Hokland P, Kassem M. Maintenance of differentiation potential of human bone marrow mesenchymal stem cells immortalized by human telomerase reverse transcriptase gene in despite of extensive proliferation. Biochem Biophys Res Commun. 2005;326:527–538. doi: 10.1016/j.bbrc.2004.11.059. - DOI - PubMed
-
- Aerts JL, Gonzales MI, Topalian SL. Selection of appropriate control genes to assess expression of tumor antigens using real-time RT-PCR. BioTechniques. 2004;36:84–91. - PubMed
-
- Andersen CL, Jensen JL, Ørntoft TF. Normalization of real-time quantitative reverse transcription-pcr data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets cancer research. Cancer Res. 2004;64:5245–5250. doi: 10.1158/0008-5472.CAN-04-0496. - DOI - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

