Sequence-specific recognition of methylated DNA by human zinc-finger proteins
- PMID: 20403812
- PMCID: PMC2926618
- DOI: 10.1093/nar/gkq280
Sequence-specific recognition of methylated DNA by human zinc-finger proteins
Abstract
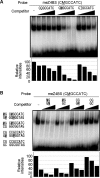
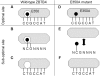
DNA methylation is an essential epigenetic mark. Three classes of mammalian proteins recognize methylated DNA: MBD proteins, SRA proteins and the zinc-finger proteins Kaiso, ZBTB4 and ZBTB38. The last three proteins can bind either methylated DNA or unmethylated consensus sequences; how this is achieved is largely unclear. Here, we report that the human zinc-finger proteins Kaiso, ZBTB4 and ZBTB38 can bind methylated DNA in a sequence-specific manner, and that they may use a mode of binding common to other zinc-finger proteins. This suggests that many other sequence-specific methyl binding proteins may exist.
Figures



Similar articles
-
Context-dependent CpG methylation directs cell-specific binding of transcription factor ZBTB38.Epigenetics. 2022 Dec;17(13):2122-2143. doi: 10.1080/15592294.2022.2111135. Epub 2022 Aug 24. Epigenetics. 2022. PMID: 36000449 Free PMC article.
-
A family of human zinc finger proteins that bind methylated DNA and repress transcription.Mol Cell Biol. 2006 Jan;26(1):169-81. doi: 10.1128/MCB.26.1.169-181.2006. Mol Cell Biol. 2006. PMID: 16354688 Free PMC article.
-
On how mammalian transcription factors recognize methylated DNA.Epigenetics. 2013 Feb;8(2):131-7. doi: 10.4161/epi.23632. Epub 2013 Jan 16. Epigenetics. 2013. PMID: 23324617 Free PMC article. Review.
-
Structural insights into methylated DNA recognition by the C-terminal zinc fingers of the DNA reader protein ZBTB38.J Biol Chem. 2018 Dec 21;293(51):19835-19843. doi: 10.1074/jbc.RA118.005147. Epub 2018 Oct 24. J Biol Chem. 2018. PMID: 30355731 Free PMC article.
-
A common mode of recognition for methylated CpG.Trends Biochem Sci. 2013 Apr;38(4):177-83. doi: 10.1016/j.tibs.2012.12.005. Epub 2013 Jan 23. Trends Biochem Sci. 2013. PMID: 23352388 Free PMC article. Review.
Cited by
-
Context-dependent CpG methylation directs cell-specific binding of transcription factor ZBTB38.Epigenetics. 2022 Dec;17(13):2122-2143. doi: 10.1080/15592294.2022.2111135. Epub 2022 Aug 24. Epigenetics. 2022. PMID: 36000449 Free PMC article.
-
The SET-domain protein SUVR5 mediates H3K9me2 deposition and silencing at stimulus response genes in a DNA methylation-independent manner.PLoS Genet. 2012;8(10):e1002995. doi: 10.1371/journal.pgen.1002995. Epub 2012 Oct 11. PLoS Genet. 2012. PMID: 23071452 Free PMC article.
-
The Mechanisms of Generation, Recognition, and Erasure of DNA 5-Methylcytosine and Thymine Oxidations.J Biol Chem. 2015 Aug 21;290(34):20723-20733. doi: 10.1074/jbc.R115.656884. Epub 2015 Jul 7. J Biol Chem. 2015. PMID: 26152719 Free PMC article. Review.
-
Depletion of ZBTB38 potentiates the effects of DNA demethylating agents in cancer cells via CDKN1C mRNA up-regulation.Oncogenesis. 2018 Oct 11;7(10):82. doi: 10.1038/s41389-018-0092-0. Oncogenesis. 2018. PMID: 30310057 Free PMC article.
-
CH···O Hydrogen Bonds Mediate Highly Specific Recognition of Methylated CpG Sites by the Zinc Finger Protein Kaiso.Biochemistry. 2018 Apr 10;57(14):2109-2120. doi: 10.1021/acs.biochem.8b00065. Epub 2018 Mar 26. Biochemistry. 2018. PMID: 29546986 Free PMC article.
References
-
- Klose RJ, Bird AP. Genomic DNA methylation: the mark and its mediators. Trends Biochem. Sci. 2006;31:89–97. - PubMed
-
- Sasai N, Defossez PA. Many paths to one goal? The proteins that recognize methylated DNA in eukaryotes. Int. J. Dev. Biol. 2009;53:323–334. - PubMed
-
- Lopes EC, Valls E, Figueroa ME, Mazur A, Meng FG, Chiosis G, Laird PW, Schreiber-Agus N, Greally JM, Prokhortchouk E, et al. Kaiso contributes to DNA methylation-dependent silencing of tumor suppressor genes in colon cancer cell lines. Cancer Res. 2008;68:7258–7263. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

