GEMIN2 promotes accumulation of RAD51 at double-strand breaks in homologous recombination
- PMID: 20403813
- PMCID: PMC2926616
- DOI: 10.1093/nar/gkq271
GEMIN2 promotes accumulation of RAD51 at double-strand breaks in homologous recombination
Abstract
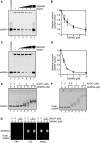
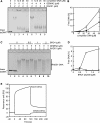
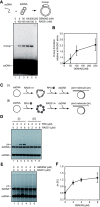
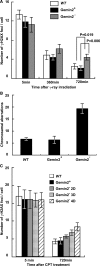
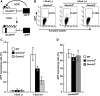
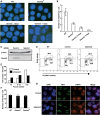
RAD51 is a key factor in homologous recombination (HR) and plays an essential role in cellular proliferation by repairing DNA damage during replication. The assembly of RAD51 at DNA damage is strictly controlled by RAD51 mediators, including BRCA1 and BRCA2. We found that human RAD51 directly binds GEMIN2/SIP1, a protein involved in spliceosome biogenesis. Biochemical analyses indicated that GEMIN2 enhances the RAD51-DNA complex formation by inhibiting RAD51 dissociation from DNA, and thereby stimulates RAD51-mediated homologous pairing. GEMIN2 also enhanced the RAD51-mediated strand exchange, when RPA was pre-bound to ssDNA before the addition of RAD51. To analyze the function of GEMIN2, we depleted GEMIN2 in the chicken DT40 line and in human cells. The loss of GEMIN2 reduced HR efficiency and resulted in a significant decrease in the number of RAD51 subnuclear foci, as observed in cells deficient in BRCA1 and BRCA2. These observations and our biochemical analyses reveal that GEMIN2 regulates HR as a novel RAD51 mediator.
Figures
Similar articles
-
Purification of the human SMN-GEMIN2 complex and assessment of its stimulation of RAD51-mediated DNA recombination reactions.Biochemistry. 2011 Aug 16;50(32):6797-805. doi: 10.1021/bi200828g. Epub 2011 Jul 25. Biochemistry. 2011. PMID: 21732698
-
BRCA1 regulates RAD51 function in response to DNA damage and suppresses spontaneous sister chromatid replication slippage: implications for sister chromatid cohesion, genome stability, and carcinogenesis.Cancer Res. 2005 Dec 15;65(24):11384-91. doi: 10.1158/0008-5472.CAN-05-2156. Cancer Res. 2005. PMID: 16357146
-
DNA Strand Exchange to Monitor Human RAD51-Mediated Strand Invasion and Pairing.Methods Mol Biol. 2021;2153:101-113. doi: 10.1007/978-1-0716-0644-5_8. Methods Mol Biol. 2021. PMID: 32840775 Free PMC article.
-
Non-Recombinogenic Functions of Rad51, BRCA2, and Rad52 in DNA Damage Tolerance.Genes (Basel). 2021 Sep 29;12(10):1550. doi: 10.3390/genes12101550. Genes (Basel). 2021. PMID: 34680945 Free PMC article. Review.
-
Therapeutic exploitation of tumor cell defects in homologous recombination.Anticancer Agents Med Chem. 2008 May;8(4):448-60. doi: 10.2174/187152008784220267. Anticancer Agents Med Chem. 2008. PMID: 18473729 Review.
Cited by
-
RAD18- and BRCA1-dependent pathways promote cellular tolerance to the nucleoside analog ganciclovir.Genes Cells. 2024 Nov;29(11):935-950. doi: 10.1111/gtc.13155. Epub 2024 Aug 22. Genes Cells. 2024. PMID: 39169841 Free PMC article.
-
DNA Damage Response and DNA Repair in Skeletal Myocytes From a Mouse Model of Spinal Muscular Atrophy.J Neuropathol Exp Neurol. 2016 Sep;75(9):889-902. doi: 10.1093/jnen/nlw064. Epub 2016 Jul 24. J Neuropathol Exp Neurol. 2016. PMID: 27452406 Free PMC article.
-
NSC Physiological Features in Spinal Muscular Atrophy: SMN Deficiency Effects on Neurogenesis.Int J Mol Sci. 2022 Dec 2;23(23):15209. doi: 10.3390/ijms232315209. Int J Mol Sci. 2022. PMID: 36499528 Free PMC article. Review.
-
The splicing-factor related protein SFPQ/PSF interacts with RAD51D and is necessary for homology-directed repair and sister chromatid cohesion.Nucleic Acids Res. 2011 Jan;39(1):132-45. doi: 10.1093/nar/gkq738. Epub 2010 Sep 2. Nucleic Acids Res. 2011. PMID: 20813759 Free PMC article.
-
Total Synthesis, Biological Evaluation, and Target Identification of Rare Abies Sesquiterpenoids.J Am Chem Soc. 2018 Dec 19;140(50):17465-17473. doi: 10.1021/jacs.8b07652. Epub 2018 Dec 6. J Am Chem Soc. 2018. PMID: 30461272 Free PMC article.
References
-
- Pommier Y. Topoisomerase I inhibitors: camptothecins and beyond. Nat. Rev. Cancer. 2006;6:789–802. - PubMed
-
- Sung PA, Libura J, Richardson C. Etoposide and illegitimate DNA double-strand break repair in the generation of MLL translocations: new insights and new questions. DNA Repair. 2006;5:1109–1118. - PubMed
-
- Sonoda E, Hochegger H, Saberi A, Taniguchi Y, Takeda S. Differential usage of non-homologous end-joining and homologous recombination in double strand break repair. DNA Repair. 2006;5:1021–1029. - PubMed
-
- Takeda S, Nakamura K, Taniguchi Y, Paull TT. Ctp1/CtIP and the MRN complex collaborate in the initial steps of homologous recombination. Mol. Cell. 2007;28:351–352. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

