A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus
- PMID: 20404139
- PMCID: PMC2889581
- DOI: 10.1073/pnas.0913535107
A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus
Abstract
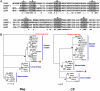
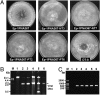
Mycoviruses are viruses that infect fungi and have the potential to control fungal diseases of crops when associated with hypovirulence. Typically, mycoviruses have double-stranded (ds) or single-stranded (ss) RNA genomes. No mycoviruses with DNA genomes have previously been reported. Here, we describe a hypovirulence-associated circular ssDNA mycovirus from the plant pathogenic fungus Sclerotinia sclerotiorum. The genome of this ssDNA virus, named Sclerotinia sclerotiorum hypovirulence-associated DNA virus 1 (SsHADV-1), is 2166 nt, coding for a replication initiation protein (Rep) and a coat protein (CP). Although phylogenetic analysis of Rep showed that SsHADV-1 is related to geminiviruses, it is notably distinct from geminiviruses both in genome organization and particle morphology. Polyethylene glycol-mediated transfection of fungal protoplasts was successful with either purified SsHADV-1 particles or viral DNA isolated directly from infected mycelium. The discovery of an ssDNA mycovirus enhances the potential of exploring fungal viruses as valuable tools for molecular manipulation of fungi and for plant disease control and expands our knowledge of global virus ecology and evolution.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
Extracellular transmission of a DNA mycovirus and its use as a natural fungicide.Proc Natl Acad Sci U S A. 2013 Jan 22;110(4):1452-7. doi: 10.1073/pnas.1213755110. Epub 2013 Jan 7. Proc Natl Acad Sci U S A. 2013. PMID: 23297222 Free PMC article.
-
Identification of the Viral Determinant of Hypovirulence and Host Range in Sclerotiniaceae of a Genomovirus Reconstructed from the Plant Metagenome.J Virol. 2021 Aug 10;95(17):e0026421. doi: 10.1128/JVI.00264-21. Epub 2021 Aug 10. J Virol. 2021. PMID: 34132570 Free PMC article.
-
A Novel RNA Virus Related to Sobemoviruses Confers Hypovirulence on the Phytopathogenic Fungus Sclerotinia sclerotiorum.Viruses. 2019 Aug 16;11(8):759. doi: 10.3390/v11080759. Viruses. 2019. PMID: 31426425 Free PMC article.
-
Viruses of the plant pathogenic fungus Sclerotinia sclerotiorum.Adv Virus Res. 2013;86:215-48. doi: 10.1016/B978-0-12-394315-6.00008-8. Adv Virus Res. 2013. PMID: 23498908 Review.
-
Mycovirus-induced hypovirulence in notorious fungi Sclerotinia: a comprehensive review.Braz J Microbiol. 2023 Sep;54(3):1459-1478. doi: 10.1007/s42770-023-01073-4. Epub 2023 Jul 31. Braz J Microbiol. 2023. PMID: 37523037 Free PMC article. Review.
Cited by
-
Novel plant-associated genomoviruses from the Brazilian Cerrado biome.Arch Virol. 2023 Nov 9;168(12):286. doi: 10.1007/s00705-023-05892-6. Arch Virol. 2023. PMID: 37940763
-
Identification and analysis of MKK and MPK gene families in canola (Brassica napus L.).BMC Genomics. 2013 Jun 11;14:392. doi: 10.1186/1471-2164-14-392. BMC Genomics. 2013. PMID: 23758924 Free PMC article.
-
The HEX1 gene of Fusarium graminearum is required for fungal asexual reproduction and pathogenesis and for efficient viral RNA accumulation of Fusarium graminearum virus 1.J Virol. 2013 Sep;87(18):10356-67. doi: 10.1128/JVI.01026-13. Epub 2013 Jul 17. J Virol. 2013. PMID: 23864619 Free PMC article.
-
A novel hypovirulence-associated Hadaka virus 1 (HadV1-LA6) in Fusarium oxysporum f. sp. cubense.mSphere. 2024 Aug 28;9(8):e0042824. doi: 10.1128/msphere.00428-24. Epub 2024 Jul 16. mSphere. 2024. PMID: 39012104 Free PMC article.
-
Characterization of a Novel Megabirnavirus from Sclerotinia sclerotiorum Reveals Horizontal Gene Transfer from Single-Stranded RNA Virus to Double-Stranded RNA Virus.J Virol. 2015 Aug;89(16):8567-79. doi: 10.1128/JVI.00243-15. Epub 2015 Jun 10. J Virol. 2015. PMID: 26063429 Free PMC article.
References
-
- Stokstad E. Plant pathology. Deadly wheat fungus threatens world's breadbaskets. Science. 2007;315:1786–1787. - PubMed
-
- Nuss DL. Hypovirulence: Mycoviruses at the fungal-plant interface. Nat Rev Microbiol. 2005;3:632–642. - PubMed
-
- Buck KW, Brasier CM. In: Viruses of the Dutch Elm Disease Fungi, in dsRNA Genetic Elements: Concepts and Applications in Agriculture, Forestry, and Medicine. Tavantzis SM, editor. Boca Raton, FL: CRC Press; 2002. pp. 165–190.
-
- Anagnostakis SL. Biological control of chestnut blight. Science. 1982;215:466–471. - PubMed
-
- Heiniger U, Rigling D. Biological control of chestnut blight in Europe. Annu Rev Phytopathol. 1994;32:581–599.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

