Measurement of protein synthesis using heavy water labeling and peptide mass spectrometry: Discrimination between major histocompatibility complex allotypes
- PMID: 20406617
- PMCID: PMC2896473
- DOI: 10.1016/j.ab.2010.04.018
Measurement of protein synthesis using heavy water labeling and peptide mass spectrometry: Discrimination between major histocompatibility complex allotypes
Abstract
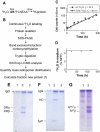
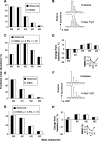
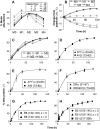
Methodological limitations have hampered the use of heavy water ((2)H(2)O), a convenient, universal biosynthetic label, for measuring protein synthesis. Analyses of (2)H-labeled amino acids are sensitive to contamination; labeling of peptides has been measured for a few serum proteins, but this approach awaits full validation. Here we describe a method for quantifying protein synthesis by peptide mass spectrometry (MS) after (2)H(2)O labeling, as applied to various proteins of the major histocompatibility complex (MHC). Human and murine antigen-presenting cells were cultured in medium containing 5% (2)H(2)O; class I and class II MHC proteins were immunoprecipitated, bands were excised, and Ala-/Gly-rich, allele-specific tryptic peptides were identified by liquid chromatography-tandem mass spectrometry (LC-MS/MS). Mass isotopomer distributions were quantified precisely by LC-MS and shifted markedly on (2)H(2)O labeling. Experimental data agreed closely with models obtained by mass isotopomer distribution analysis (MIDA) and were consistent with contributions from Ala, Gly, and other amino acids to labeling. Estimates of fractional protein synthesis from peptides of the same protein were precise and internally consistent. The method was capable of discriminating between MHC isotypes and alleles, applicable to primary cells, and readily extendable to other proteins. It simplifies measurements of protein synthesis, enabling novel applications in physiology, in genotype/phenotype interactions, and potentially in kinetic proteomics.
Copyright 2010 Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Measurement of protein turnover rates by heavy water labeling of nonessential amino acids.Biochim Biophys Acta. 2006 May;1760(5):730-44. doi: 10.1016/j.bbagen.2005.12.023. Epub 2006 Jan 24. Biochim Biophys Acta. 2006. PMID: 16567052
-
Measuring protein synthesis by mass isotopomer distribution analysis (MIDA).Anal Biochem. 1999 Feb 1;267(1):1-16. doi: 10.1006/abio.1998.2958. Anal Biochem. 1999. PMID: 9918649
-
Protein turnover models for LC-MS data of heavy water metabolic labeling.Brief Bioinform. 2022 Mar 10;23(2):bbab598. doi: 10.1093/bib/bbab598. Brief Bioinform. 2022. PMID: 35062023 Free PMC article.
-
Dynamic Proteomics: In Vivo Proteome-Wide Measurement of Protein Kinetics Using Metabolic Labeling.Methods Enzymol. 2015;561:219-76. doi: 10.1016/bs.mie.2015.05.018. Epub 2015 Jul 17. Methods Enzymol. 2015. PMID: 26358907 Review.
-
Mass isotopomer distribution analysis at eight years: theoretical, analytic, and experimental considerations.Am J Physiol. 1999 Jun;276(6):E1146-70. doi: 10.1152/ajpendo.1999.276.6.E1146. Am J Physiol. 1999. PMID: 10362629 Review.
Cited by
-
Metabolic labeling reveals proteome dynamics of mouse mitochondria.Mol Cell Proteomics. 2012 Dec;11(12):1586-94. doi: 10.1074/mcp.M112.021162. Epub 2012 Aug 21. Mol Cell Proteomics. 2012. PMID: 22915825 Free PMC article.
-
In vivo deuterium magnetic resonance imaging of xenografted tumors following systemic administration of deuterated water.Sci Rep. 2023 Sep 7;13(1):14699. doi: 10.1038/s41598-023-41163-9. Sci Rep. 2023. PMID: 37679461 Free PMC article.
-
The effect of long term calorie restriction on in vivo hepatic proteostatis: a novel combination of dynamic and quantitative proteomics.Mol Cell Proteomics. 2012 Dec;11(12):1801-14. doi: 10.1074/mcp.M112.021204. Epub 2012 Sep 13. Mol Cell Proteomics. 2012. PMID: 22984287 Free PMC article.
-
Isotope-Substitution Effects on the Thermodynamic, Dynamic, and Structural Properties of Water: H2O, HDO, D2O, and T2O.J Phys Chem B. 2025 Jul 10;129(27):6886-6902. doi: 10.1021/acs.jpcb.5c01657. Epub 2025 Jun 29. J Phys Chem B. 2025. PMID: 40581977 Free PMC article.
-
Exploring ribosome composition and newly synthesized proteins through proteomics and potential biomedical applications.Expert Rev Proteomics. 2017 Jun;14(6):529-543. doi: 10.1080/14789450.2017.1333424. Epub 2017 May 26. Expert Rev Proteomics. 2017. PMID: 28532181 Free PMC article. Review.
References
-
- Papageorgopoulos C., Caldwell K., Shackleton C., Schweingrubber H., Hellerstein M.K. Measuring protein synthesis by mass isotopomer distribution analysis (MIDA) Anal. Biochem. 1999;267:1–16. - PubMed
-
- Kruger M., Moser M., Ussar S., Thievessen I., Luber C.A., Forner F., Schmidt S., Zanivan S., Fassler R., Mann M. SILAC mouse for quantitative proteomics uncovers kindlin-3 as an essential factor for red blood cell function. Cell. 2008;134:353–364. - PubMed
-
- Harsha H.C., Molina H., Pandey A. Quantitative proteomics using stable isotope labeling with amino acids in cell culture. Nat. Protoc. 2008;3:505–516. - PubMed
-
- Beynon R.J., Pratt J.M. Metabolic labeling of proteins for proteomics. Mol. Cell. Proteomics. 2005;4:857–872. - PubMed
-
- Dufner D.A., Bederman I.R., Brunengraber D.Z., Rachdaoui N., Ismail-Beigi F., Siegfried B.A., Kimball S.R., Previs S.F. Using 2H2O to study the influence of feeding on protein synthesis: effect of isotope equilibration in vivo vs. in cell culture. Am. J. Physiol. Endocrinol. Metab. 2005;288:E1277–E1283. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

