Mitotic expression of Spo13 alters M-phase progression and nucleolar localization of Cdc14 in budding yeast
- PMID: 20407133
- PMCID: PMC2897760
- DOI: 10.1534/genetics.109.113746
Mitotic expression of Spo13 alters M-phase progression and nucleolar localization of Cdc14 in budding yeast
Abstract
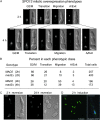
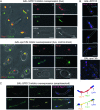
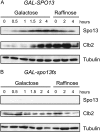
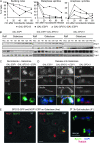
Spo13 is a key meiosis-specific regulator required for centromere cohesion and coorientation, and for progression through two nuclear divisions. We previously reported that it causes a G2/M arrest and may delay the transition from late anaphase to G1, when overexpressed in mitosis. Yet its mechanism of action has remained elusive. Here we show that Spo13, which is phosphorylated and stabilized at G2/M in a Cdk/Clb-dependent manner, acts at two stages during mitotic cell division. Spo13 provokes a G2/M arrest that is reversible and largely independent of the Mad2 spindle checkpoint. Since mRNAs whose induction requires Cdc14 activation are reduced, we propose that its anaphase delay results from inhibition of Cdc14 function. Indeed, the Spo13-induced anaphase delay correlates with Cdc14 phosphatase retention in the nucleolus and with cyclin B accumulation, which both impede anaphase exit. At the onset of arrest, Spo13 is primarily associated with the nucleolus, where Cdc14 accumulates. Significantly, overexpression of separase (Esp1), which promotes G2/M and anaphase progression, suppresses Spo13 effects in mitosis, arguing that Spo13 acts upstream or parallel to Esp1. Given that Spo13 overexpression reduces Pds1 and cyclin B degradation, our findings are consistent with a role for Spo13 in regulating APC, which controls both G2/M and anaphase. Similar effects of Spo13 during meiotic MI may prevent cell cycle exit and initiation of DNA replication prior to MII, thereby ensuring two successive chromosome segregation events without an intervening S phase.
Figures


Similar articles
-
Phosphorylation by cyclin B-Cdk underlies release of mitotic exit activator Cdc14 from the nucleolus.Science. 2004 Jul 23;305(5683):516-9. doi: 10.1126/science.1099402. Science. 2004. PMID: 15273393
-
Mitotic exit in two dimensions.J Theor Biol. 2007 Oct 7;248(3):560-73. doi: 10.1016/j.jtbi.2007.06.014. Epub 2007 Jun 17. J Theor Biol. 2007. PMID: 17659305
-
Cdc14 Early Anaphase Release, FEAR, Is Limited to the Nucleus and Dispensable for Efficient Mitotic Exit.PLoS One. 2015 Jun 19;10(6):e0128604. doi: 10.1371/journal.pone.0128604. eCollection 2015. PLoS One. 2015. PMID: 26090959 Free PMC article.
-
The Multiple Roles of the Cdc14 Phosphatase in Cell Cycle Control.Int J Mol Sci. 2020 Jan 21;21(3):709. doi: 10.3390/ijms21030709. Int J Mol Sci. 2020. PMID: 31973188 Free PMC article. Review.
-
Decoding the Nucleolar Role in Meiotic Recombination and Cell Cycle Control: Insights into Cdc14 Function.Int J Mol Sci. 2024 Nov 29;25(23):12861. doi: 10.3390/ijms252312861. Int J Mol Sci. 2024. PMID: 39684572 Free PMC article. Review.
Cited by
-
Global alterations of the transcriptional landscape during yeast growth and development in the absence of Ume6-dependent chromatin modification.Mol Genet Genomics. 2015 Oct;290(5):2031-46. doi: 10.1007/s00438-015-1051-5. Epub 2015 May 10. Mol Genet Genomics. 2015. PMID: 25957495 Free PMC article.
-
Key factors influencing undergraduate nursing students' perceptions of the use of learning management systems: a systematic literature review.BMC Nurs. 2025 Mar 26;24(1):323. doi: 10.1186/s12912-025-02962-9. BMC Nurs. 2025. PMID: 40140804 Free PMC article.
-
Construction of strains to identify novel factors for regulation of centromeric cohesion protection (CCP) and sister kinetochore mono-orientation (SKM).BMC Mol Cell Biol. 2019 Oct 22;20(1):44. doi: 10.1186/s12860-019-0231-2. BMC Mol Cell Biol. 2019. PMID: 31640543 Free PMC article.
References
-
- Azzam, R., S. L. Chen, W. Shou, G. Alexandru, K. Nasmyth et al., 2004. Phosphorylation by cyclin B-Cdk underlies release of mitotic exit activator Cdc14 from the nucleolus. Science 305 516–519. - PubMed
-
- Bishop, A. C., J. A. Ubersax, D. T. Petsch, D. P. Matheos, N. S. Gray et al., 2000. A chemical switch for inhibitor-sensitive alleles of any protein kinase. Nature 407 395–401. - PubMed
-
- Brown, N. R., M. E. Noble, J. A. Endicott and L. N. Johnson, 1999. The structural basis for specificity of substrate and recruitment peptides for cyclin-dependent kinases. Nat. Cell Biol. 1 438–443. - PubMed
-
- Buonomo, S. B., K. P. Rabitsch, J. Fuchs, S. Gruber, M. Sullivan et al., 2003. Division of the nucleolus and its release of CDC14 during anaphase of meiosis I depends on separase, SPO12, and SLK19. Dev. Cell 4 727–739. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

