S-layer stabilized lipid membranes (Review)
- PMID: 20408666
- PMCID: PMC2876325
- DOI: 10.1116/1.2889067
S-layer stabilized lipid membranes (Review)
Abstract
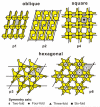
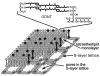
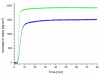
The present review focuses on a unique bio-molecular construction kit based on surface-layer (S-layer) proteins as building blocks and patterning elements, but also major classes of biological molecules such as lipids, membrane-active peptides and membrane proteins, and glycans for the design of functional supported lipid membranes. The biomimetic approach copying the supramolecular building principle of most archaeal cell envelopes merely composed of a plasma membrane and a closely associated S-layer lattice has resulted in robust and fluid lipid membranes. Most importantly, S-layer supported lipid membranes spanning an aperture or generated on solid and porous substrates constitute highly interesting model membranes for the reconstitution of responsive transmembrane proteins and membrane-active peptides. This is of particular challenge as one-third of all proteins are membrane proteins such as pore-forming proteins, ion channels, and receptors. S-layer supported lipid membranes are seen as one of the most innovative strategies in membrane protein-based nanobiotechnology with potential applications that range from pharmaceutical (high-throughput) drug screening over lipid chips to the detection of biological warfare agents.
Figures

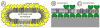


Similar articles
-
Biomimetic interfaces based on S-layer proteins, lipid membranes and functional biomolecules.J R Soc Interface. 2014 May 8;11(96):20140232. doi: 10.1098/rsif.2014.0232. Print 2014 Jul 6. J R Soc Interface. 2014. PMID: 24812051 Free PMC article. Review.
-
Composite S-layer lipid structures.J Struct Biol. 2009 Oct;168(1):207-16. doi: 10.1016/j.jsb.2009.03.004. Epub 2009 Mar 20. J Struct Biol. 2009. PMID: 19303933 Free PMC article. Review.
-
Solid supported lipid membranes: new concepts for the biomimetic functionalization of solid surfaces.Biointerphases. 2008 Jun;3(2):FA125. doi: 10.1116/1.2913612. Biointerphases. 2008. PMID: 20408662 Free PMC article.
-
S-layers as a tool kit for nanobiotechnological applications.FEMS Microbiol Lett. 2007 Feb;267(2):131-44. doi: 10.1111/j.1574-6968.2006.00573.x. FEMS Microbiol Lett. 2007. PMID: 17328112 Review.
-
Nanobiotechnology with S-layer proteins as building blocks.Prog Mol Biol Transl Sci. 2011;103:277-352. doi: 10.1016/B978-0-12-415906-8.00003-0. Prog Mol Biol Transl Sci. 2011. PMID: 21999999 Review.
Cited by
-
S-layer protein self-assembly.Int J Mol Sci. 2013 Jan 25;14(2):2484-501. doi: 10.3390/ijms14022484. Int J Mol Sci. 2013. PMID: 23354479 Free PMC article.
-
Relevance of glycosylation of S-layer proteins for cell surface properties.Acta Biomater. 2015 Jun;19:149-157. doi: 10.1016/j.actbio.2015.03.020. Epub 2015 Mar 25. Acta Biomater. 2015. PMID: 25818946 Free PMC article.
-
Electrochemical Biosensors Based on S-Layer Proteins.Sensors (Basel). 2020 Mar 19;20(6):1721. doi: 10.3390/s20061721. Sensors (Basel). 2020. PMID: 32204503 Free PMC article. Review.
-
Assorted Methods for Decontamination of Aflatoxin M1 in Milk Using Microbial Adsorbents.Toxins (Basel). 2019 May 29;11(6):304. doi: 10.3390/toxins11060304. Toxins (Basel). 2019. PMID: 31146398 Free PMC article. Review.
-
Development of functionalised polyelectrolyte capsules using filamentous Escherichia coli cells.Microb Cell Fact. 2012 Dec 23;11:163. doi: 10.1186/1475-2859-11-163. Microb Cell Fact. 2012. PMID: 23259586 Free PMC article.
References
-
- Gerstein M, Hegyi H. FEMS Microbiol. Rev. 1998;22:277. - PubMed
-
- Galdiero S, Galdiero M, Pedone C. Curr. Protein Pept. Sci. 2007;8:63. - PubMed
-
- Viviani B, Gardoni F, Marinovich M. Int. Rev. Neurobiol. 2007;82:247. - PubMed
-
- Ellis C, Smith A. Nat. Rev. Drug Discovery. 2004;3:237. - PubMed
-
- Henrick K, Feng Z, Bluhm WF, Dimitropoulos D, Doreleijers JF, Dutta S, Flippen-Anderson JL, Ionides J, Kamada C, Krissinel E, Lawson CL, Markley JL, Nakamura H, Newman R, Shimizu Y, Swaminathan J, Velankar S, Ory J, Ulrich EL, Vranken W, Westbrook J, Yamashita R, Yang H, Young J, Yousufuddin M, Berman HM. Nucleic Acids Res. 2007;36:D426. - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

