Identification and characterization of putative methylation targets in the MAOA locus using bioinformatic approaches
- PMID: 20421737
- PMCID: PMC3169210
- DOI: 10.4161/epi.5.4.11719
Identification and characterization of putative methylation targets in the MAOA locus using bioinformatic approaches
Abstract
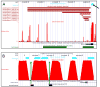
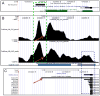
Monoamine oxidase A (MAO A) is an enzyme that catalyzes the oxidation of neurotransmitter amines. A functional polymorphism in the human MAOA gene (high- and low-MAOA) has been associated with distinct behavioral phenotypes. To investigate directly the biological mechanism whereby this polymorphism influences brain function, we recently measured the activity of the MAO A enzyme in healthy volunteers. When found no relationship between the individual's brain MAO A level and the MAOA genotype, we postulated that there are additional regulatory mechanisms that control the MAOA expression. Given that DNA methylation is linked to the regulation of gene expression, we hypothesized that epigenetic mechanisms factor into the MAOA expression. Our underplaying assumption was that the differences in an individual's genotype play a key role in the epigenetic potential of the MAOA locus and, consequently, determine the individual's level of MAO A activity in the brain. As a first step towards experimental validation of the hypothesis, we performed a comprehensive bioinformatic analysis aiming to interrogate genomic features and attributes of the MAOA locus that might modulate its epigenetic sensitivity. Major findings of our analysis are the following: (1) the extended MAOA regulatory region contains two CpG islands (CGIs), one of which overlaps with the canonical MAOA promoter and the other is located further upstream; both CGIs exhibit sensitivity to differential methylation. (2) The uVNTR's effect on the MAOA's transcriptional activity might have epigenetic nature: this polymorphic region resides within the MAOA's CGI and itself contains CpGs, thus, the number of repeating increments effectively changes the number of methylatable cytosines in the MAOA promoter. An array of in silico analyses (the nucleosome positioning, the physical properties of the local DNA, the clustering of transcription-factor binding sites) together with experimental data on histone modifications and Pol 2 sites and data from the RefSeq mRNA library suggest that the MAOA gene might have an alternative promoter. Based on our findings, we propose a regulatory mechanism for the human MAOA according to which the MAOA expression in vivo is executed by the generation of tissue-specific transcripts initiated from the alternative promoters (both CGI-associated) where transcriptional activation of a particular promoter is under epigenetic control.
Conflict of interest statement
Figures






Similar articles
-
Evidence that the methylation state of the monoamine oxidase A (MAOA) gene predicts brain activity of MAO A enzyme in healthy men.Epigenetics. 2012 Oct;7(10):1151-60. doi: 10.4161/epi.21976. Epub 2012 Sep 4. Epigenetics. 2012. PMID: 22948232 Free PMC article.
-
Allelic mRNA expression of X-linked monoamine oxidase a (MAOA) in human brain: dissection of epigenetic and genetic factors.Hum Mol Genet. 2006 Sep 1;15(17):2636-49. doi: 10.1093/hmg/ddl192. Epub 2006 Aug 7. Hum Mol Genet. 2006. PMID: 16893905
-
Association of DNA methylation and monoamine oxidase A gene expression in the brains of different dog breeds.Gene. 2016 Apr 15;580(2):177-182. doi: 10.1016/j.gene.2016.01.022. Epub 2016 Jan 16. Gene. 2016. PMID: 26784655
-
Understanding the interplay between CpG island-associated gene promoters and H3K4 methylation.Biochim Biophys Acta Gene Regul Mech. 2020 Aug;1863(8):194567. doi: 10.1016/j.bbagrm.2020.194567. Epub 2020 Apr 29. Biochim Biophys Acta Gene Regul Mech. 2020. PMID: 32360393 Free PMC article. Review.
-
Sequence determinants, function, and evolution of CpG islands.Biochem Soc Trans. 2021 Jun 30;49(3):1109-1119. doi: 10.1042/BST20200695. Biochem Soc Trans. 2021. PMID: 34156435 Free PMC article. Review.
Cited by
-
Evidence that the methylation state of the monoamine oxidase A (MAOA) gene predicts brain activity of MAO A enzyme in healthy men.Epigenetics. 2012 Oct;7(10):1151-60. doi: 10.4161/epi.21976. Epub 2012 Sep 4. Epigenetics. 2012. PMID: 22948232 Free PMC article.
-
Gene x disease interaction on orbitofrontal gray matter in cocaine addiction.Arch Gen Psychiatry. 2011 Mar;68(3):283-94. doi: 10.1001/archgenpsychiatry.2011.10. Arch Gen Psychiatry. 2011. PMID: 21383264 Free PMC article.
-
Serotonin Transporter Binding Potentials in Brain of Juvenile Monkeys 1 Year After Discontinuation of a 2-Year Treatment With Fluoxetine.Biol Psychiatry Cogn Neurosci Neuroimaging. 2019 Nov;4(11):948-955. doi: 10.1016/j.bpsc.2019.06.012. Epub 2019 Jul 6. Biol Psychiatry Cogn Neurosci Neuroimaging. 2019. PMID: 31471184 Free PMC article.
-
Monoamine oxidase A expression is suppressed in human cholangiocarcinoma via coordinated epigenetic and IL-6-driven events.Lab Invest. 2012 Oct;92(10):1451-60. doi: 10.1038/labinvest.2012.110. Epub 2012 Aug 20. Lab Invest. 2012. PMID: 22906985 Free PMC article.
-
Epigenetic signature of MAOA and MAOB genes in mental disorders.J Neural Transm (Vienna). 2018 Nov;125(11):1581-1588. doi: 10.1007/s00702-018-1929-6. Epub 2018 Sep 21. J Neural Transm (Vienna). 2018. PMID: 30242487 Review.
References
-
- Stahl SM, Felker A. Monoamine oxidase inhibitors: a modern guide to an unrequited class of antidepressants. CNS spectrums. 2008;13:855–70. - PubMed
-
- Meyer JH, Ginovart N, Boovariwala A, Sagrati S, Hussey D, Garcia A, Young T, Praschak-Rieder N, Wilson AA, Houle S. Elevated monoamine oxidase a levels in the brain: an explanation for the monoamine imbalance of major depression. Archives of general psychiatry. 2006;63:1209–16. - PubMed
-
- Brunner HG, Nelen M, Breakefield XO, Ropers HH, van Oost BA. Abnormal behavior associated with a point mutation in the structural gene for monoamine oxidase A. Science. 1993;262:578–80. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
