Cildb: a knowledgebase for centrosomes and cilia
- PMID: 20428338
- PMCID: PMC2860946
- DOI: 10.1093/database/bap022
Cildb: a knowledgebase for centrosomes and cilia
Abstract
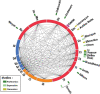
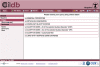
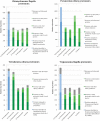
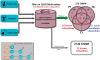
Ciliopathies, pleiotropic diseases provoked by defects in the structure or function of cilia or flagella, reflect the multiple roles of cilia during development, in stem cells, in somatic organs and germ cells. High throughput studies have revealed several hundred proteins that are involved in the composition, function or biogenesis of cilia. The corresponding genes are potential candidates for orphan ciliopathies. To study ciliary genes, model organisms are used in which particular questions on motility, sensory or developmental functions can be approached by genetics. In the course of high throughput studies of cilia in Paramecium tetraurelia, we were confronted with the problem of comparing our results with those obtained in other model organisms. We therefore developed a novel knowledgebase, Cildb, that integrates ciliary data from heterogeneous sources. Cildb links orthology relationships among 18 species to high throughput ciliary studies, and to OMIM data on human hereditary diseases. The web interface of Cildb comprises three tools, BioMart for complex queries, BLAST for sequence homology searches and GBrowse for browsing the human genome in relation to OMIM information for human diseases. Cildb can be used for interspecies comparisons, building candidate ciliary proteomes in any species, or identifying candidate ciliopathy genes.Database URL:http://cildb.cgm.cnrs-gif.fr.
Figures

References
-
- Bornens M. Organelle positioning and cell polarity. Nat. Rev. Mol. Cell Biol. 2008;9:874–886. - PubMed
-
- Dawe HR, Farr H, Gull K. Centriole/basal body morphogenesis and migration during ciliogenesis in animal cells. J. Cell Sci. 2007;120:7–15. - PubMed
-
- Marshall WF. Basal bodies platforms for building cilia. Curr. Top Dev. Biol. 2008;85:1–22. - PubMed
-
- Basu B, Brueckner M. Cilia multifunctional organelles at the center of vertebrate left-right asymmetry. Curr. Top Dev. Biol. 2008;85:151–174. - PubMed
-
- Salathe M. Regulation of mammalian ciliary beating. Annu. Rev. Physiol. 2007;69:401–22. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

