The genome sequence of the spontaneously hypertensive rat: Analysis and functional significance
- PMID: 20430781
- PMCID: PMC2877576
- DOI: 10.1101/gr.103499.109
The genome sequence of the spontaneously hypertensive rat: Analysis and functional significance
Abstract
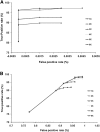
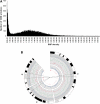
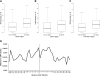
The spontaneously hypertensive rat (SHR) is the most widely studied animal model of hypertension. Scores of SHR quantitative loci (QTLs) have been mapped for hypertension and other phenotypes. We have sequenced the SHR/OlaIpcv genome at 10.7-fold coverage by paired-end sequencing on the Illumina platform. We identified 3.6 million high-quality single nucleotide polymorphisms (SNPs) between the SHR/OlaIpcv and Brown Norway (BN) reference genome, with a high rate of validation (sensitivity 96.3%-98.0% and specificity 99%-100%). We also identified 343,243 short indels between the SHR/OlaIpcv and reference genomes. These SNPs and indels resulted in 161 gain or loss of stop codons and 629 frameshifts compared with the BN reference sequence. We also identified 13,438 larger deletions that result in complete or partial absence of 107 genes in the SHR/OlaIpcv genome compared with the BN reference and 588 copy number variants (CNVs) that overlap with the gene regions of 688 genes. Genomic regions containing genes whose expression had been previously mapped as cis-regulated expression quantitative trait loci (eQTLs) were significantly enriched with SNPs, short indels, and larger deletions, suggesting that some of these variants have functional effects on gene expression. Genes that were affected by major alterations in their coding sequence were highly enriched for genes related to ion transport, transport, and plasma membrane localization, providing insights into the likely molecular and cellular basis of hypertension and other phenotypes specific to the SHR strain. This near complete catalog of genomic differences between two extensively studied rat strains provides the starting point for complete elucidation, at the molecular level, of the physiological and pathophysiological phenotypic differences between individuals from these strains.
Figures


References
-
- Agnihotri G, Liu HW 2003. Enoyl-CoA hydratase. Reaction, mechanism, and inhibition. Bioorg Med Chem 11: 9–20 - PubMed
-
- Aitman TJ, Glazier AM, Wallace CA, Cooper LD, Norsworthy PJ, Wahid FN, Al-Majali KM, Trembling PM, Mann CJ, Shoulders CC, et al. 1999. Identification of Cd36 (Fat) as an insulin-resistance gene causing defective fatty acid and glucose metabolism in hypertensive rats. Nat Genet 21: 76–83 - PubMed
-
- Aitman TJ, Critser JK, Cuppen E, Dominiczak A, Fernandez-Suarez XM, Flint J, Gauguier D, Geurts AM, Gould M, Harris PC, et al. 2008. Progress and prospects in rat genetics: A community view. Nat Genet 40: 516–522 - PubMed
-
- Andreasen D, Friis UG, Uhrenholt TR, Jensen BL, Skott O, Hansen PB 2006. Coexpression of voltage-dependent calcium channels Cav1.2, 2.1a, and 2.1b in vascular myocytes. Hypertension 47: 735–741 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
