Adaptive evolution of Escherichia coli K-12 MG1655 during growth on a Nonnative carbon source, L-1,2-propanediol
- PMID: 20435762
- PMCID: PMC2897412
- DOI: 10.1128/AEM.00373-10
Adaptive evolution of Escherichia coli K-12 MG1655 during growth on a Nonnative carbon source, L-1,2-propanediol
Erratum in
- Appl Environ Microbiol. 2010 Sep;76(18):6327
Abstract
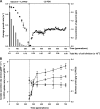
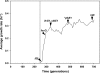
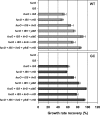
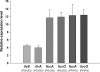
Laboratory adaptive evolution studies can provide key information to address a wide range of issues in evolutionary biology. Such studies have been limited thus far by the inability of workers to readily detect mutations in evolved microbial strains on a genome scale. This limitation has now been overcome by recently developed genome sequencing technology that allows workers to identify all accumulated mutations that appear during laboratory adaptive evolution. In this study, we evolved Escherichia coli K-12 MG1655 with a nonnative carbon source, l-1,2-propanediol (l-1,2-PDO), for approximately 700 generations. We found that (i) experimental evolution of E. coli for approximately 700 generations in 1,2-PDO-supplemented minimal medium resulted in acquisition of the ability to use l-1,2-PDO as a sole carbon and energy source so that the organism changed from an organism that did not grow at all initially to an organism that had a growth rate of 0.35 h(-1); (ii) six mutations detected by whole-genome resequencing accumulated in the evolved E. coli mutant over the course of adaptive evolution on l-1,2-PDO; (iii) five of the six mutations were within coding regions, and IS5 was inserted between two fuc regulons; (iv) two major mutations (mutations in fucO and its promoter) involved in l-1,2-PDO catabolism appeared early during adaptive evolution; and (v) multiple defined knock-in mutant strains with all of the mutations had growth rates essentially matching that of the evolved strain. These results provide insight into the genetic basis underlying microbial evolution for growth on a nonnative substrate.
Figures

References
-
- Barrick, J. E., D. S. Yu, S. H. Yoon, H. Jeong, T. K. Oh, D. Schneider, R. E. Lenski, and J. F. Kim. 2009. Genome evolution and adaptation in a long-term experiment with Escherichia coli. Nature 461:1243-1247. - PubMed
-
- Brückner, R., and F. Titgemeyer. 2002. Carbon catabolite repression in bacteria: choice of the carbon source and autoregulatory limitation of sugar utilization. FEMS Microbiol. Lett. 209:141-148. - PubMed
-
- Buckling, A., M. A. Wills, and N. Colegrave. 2003. Adaptation limits diversification of experimental bacterial populations. Science 302:2107-2109. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

