Pbx1 represses osteoblastogenesis by blocking Hoxa10-mediated recruitment of chromatin remodeling factors
- PMID: 20439491
- PMCID: PMC2897555
- DOI: 10.1128/MCB.00889-09
Pbx1 represses osteoblastogenesis by blocking Hoxa10-mediated recruitment of chromatin remodeling factors
Abstract
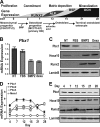
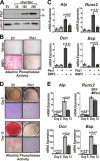
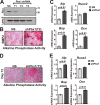
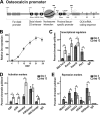
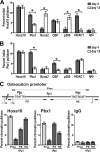
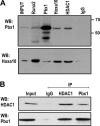
Abdominal-class homeodomain-containing (Hox) factors form multimeric complexes with TALE-class homeodomain proteins (Pbx, Meis) to regulate tissue morphogenesis and skeletal development. Here we have established that Pbx1 negatively regulates Hoxa10-mediated gene transcription in mesenchymal cells and identified components of a Pbx1 complex associated with genes in osteoblasts. Expression of Pbx1 impaired osteogenic commitment of C3H10T1/2 multipotent cells and differentiation of MC3T3-E1 preosteoblasts. Conversely, targeted depletion of Pbx1 by short hairpin RNA (shRNA) increased expression of osteoblast-related genes. Studies using wild-type and mutated osteocalcin and Bsp promoters revealed that Pbx1 acts through a Pbx-binding site that is required to attenuate gene activation by Hoxa10. Chromatin-associated Pbx1 and Hoxa10 were present at osteoblast-related gene promoters preceding gene expression, but only Hoxa10 was associated with these promoters during transcription. Our results show that Pbx1 is associated with histone deacetylases normally linked with chromatin inactivation. Loss of Pbx1 from osteoblast promoters in differentiated osteoblasts was associated with increased histone acetylation and CBP/p300 recruitment, as well as decreased H3K9 methylation. We propose that Pbx1 plays a central role in attenuating the ability of Hoxa10 to activate osteoblast-related genes in order to establish temporal regulation of gene expression during osteogenesis.
Figures



References
-
- Allen, T. D., Y. X. Zhu, T. S. Hawley, and R. G. Hawley. 2000. TALE homeoproteins as HOX11-interacting partners in T-cell leukemia. Leuk. Lymphoma 39:241-256. - PubMed
-
- Balint, E., D. Lapointe, H. Drissi, C. van der Meijden, D. W. Young, A. J. van Wijnen, J. L. Stein, G. S. Stein, and J. B. Lian. 2003. Phenotype discovery by gene expression profiling: mapping of biological processes linked to BMP-2-mediated osteoblast differentiation. J. Cell. Biochem. 89:401-426. - PubMed
-
- Bei, L., Y. Lu, S. L. Bellis, W. Zhou, E. Horvath, and E. A. Eklund. 2007. Identification of a HoxA10 activation domain necessary for transcription of the gene encoding beta3 integrin during myeloid differentiation. J. Biol. Chem. 282:16846-16859. - PubMed
-
- Berkes, C. A., D. A. Bergstrom, B. H. Penn, K. J. Seaver, P. S. Knoepfler, and S. J. Tapscott. 2004. Pbx marks genes for activation by MyoD indicating a role for a homeodomain protein in establishing myogenic potential. Mol. Cell 14:465-477. - PubMed
-
- Canalis, E., V. Deregowski, R. C. Pereira, and E. Gazzerro. 2005. Signals that determine the fate of osteoblastic cells. J. Endocrinol. Invest. 28:3-7. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous
