Predicting interaction sites from the energetics of isolated proteins: a new approach to epitope mapping
- PMID: 20441761
- PMCID: PMC2862194
- DOI: 10.1016/j.bpj.2010.01.014
Predicting interaction sites from the energetics of isolated proteins: a new approach to epitope mapping
Abstract
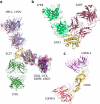
An increasing number of functional studies of proteins have shown that sequence and structural similarities alone may not be sufficient for reliable prediction of their interaction properties. This is particularly true for proteins recognizing specific antibodies, where the prediction of antibody-binding sites, called epitopes, has proven challenging. The antibody-binding properties of an antigen depend on its structure and related dynamics. Aiming to predict the antibody-binding regions of a protein, we investigate a new approach based on the integrated analysis of the dynamical and energetic properties of antigens, to identify nonoptimized, low-intensity energetic interaction networks in the protein structure isolated in solution. The method is based on the idea that recognition sites may correspond to localized regions with low-intensity energetic couplings with the rest of the protein, which allows them to undergo conformational changes, to be recognized by a binding partner, and to tolerate mutations with minimal energetic expense. Upon analyzing the results on isolated proteins and benchmarking against antibody complexes, it is found that the method successfully identifies binding sites located on the protein surface that are accessible to putative binding partners. The combination of dynamics and energetics can thus discriminate between epitopes and other substructures based only on physical properties. We discuss implications for vaccine design.
Copyright (c) 2010 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures


Similar articles
-
Rational epitope design for protein targeting.ACS Chem Biol. 2013 Feb 15;8(2):397-404. doi: 10.1021/cb300487u. Epub 2012 Nov 15. ACS Chem Biol. 2013. PMID: 23138758
-
CEP: a conformational epitope prediction server.Nucleic Acids Res. 2005 Jul 1;33(Web Server issue):W168-71. doi: 10.1093/nar/gki460. Nucleic Acids Res. 2005. PMID: 15980448 Free PMC article.
-
In Silico Prediction of Linear B-Cell Epitopes on Proteins.Methods Mol Biol. 2017;1484:255-264. doi: 10.1007/978-1-4939-6406-2_17. Methods Mol Biol. 2017. PMID: 27787831 Free PMC article.
-
Epitope mapping: the first step in developing epitope-based vaccines.BioDrugs. 2007;21(3):145-56. doi: 10.2165/00063030-200721030-00002. BioDrugs. 2007. PMID: 17516710 Free PMC article. Review.
-
Phage display for epitope determination: a paradigm for identifying receptor-ligand interactions.Biotechnol Annu Rev. 2004;10:151-88. doi: 10.1016/S1387-2656(04)10006-9. Biotechnol Annu Rev. 2004. PMID: 15504706 Review.
Cited by
-
Evaluation of docking procedures reliability in affitins-partners interactions.Front Chem. 2022 Dec 1;10:1074249. doi: 10.3389/fchem.2022.1074249. eCollection 2022. Front Chem. 2022. PMID: 36531315 Free PMC article.
-
Peptides for Infectious Diseases: From Probe Design to Diagnostic Microarrays.Antibodies (Basel). 2019 Mar 12;8(1):23. doi: 10.3390/antib8010023. Antibodies (Basel). 2019. PMID: 31544829 Free PMC article. Review.
-
Computational Analysis of Dengue Virus Envelope Protein (E) Reveals an Epitope with Flavivirus Immunodiagnostic Potential in Peptide Microarrays.Int J Mol Sci. 2019 Apr 18;20(8):1921. doi: 10.3390/ijms20081921. Int J Mol Sci. 2019. PMID: 31003530 Free PMC article.
-
Function and Potentials of M. tuberculosis Epitopes.Front Immunol. 2014 Mar 24;5:107. doi: 10.3389/fimmu.2014.00107. eCollection 2014. Front Immunol. 2014. PMID: 24715888 Free PMC article. Review.
-
SARS-CoV-2 Spike Protein Mutations and Escape from Antibodies: A Computational Model of Epitope Loss in Variants of Concern.J Chem Inf Model. 2021 Sep 27;61(9):4687-4700. doi: 10.1021/acs.jcim.1c00857. Epub 2021 Sep 1. J Chem Inf Model. 2021. PMID: 34468141 Free PMC article.
References
-
- de Vries S.J., Bonvin A.M.J.J. How proteins get in touch: interface prediction in the study of biomolecular complexes. Curr. Protein Pept. Sci. 2008;9:394–406. - PubMed
-
- Rodier F., Bahadur R.P., Janin J. Hydration of protein-protein interfaces. Proteins: Struct. Funct. Bioinf. 2005;60:36–45. - PubMed
-
- Chakrabarti P., Janin J. Dissecting protein-protein recognition sites. Proteins: Struct. Funct. Genet. 2002;47:334–343. - PubMed
-
- Keskin O., Gursoy A., Nussinov R. Principles of protein-protein interactions: what are the preferred ways for proteins to interact? Chem. Rev. 2008;108:1225–1244. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

