WRN mutations in Werner syndrome patients: genomic rearrangements, unusual intronic mutations and ethnic-specific alterations
- PMID: 20443122
- PMCID: PMC4686336
- DOI: 10.1007/s00439-010-0832-5
WRN mutations in Werner syndrome patients: genomic rearrangements, unusual intronic mutations and ethnic-specific alterations
Abstract
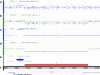
Werner syndrome (WS) is an autosomal recessive segmental progeroid syndrome caused by null mutations at the WRN locus, which codes for a member of the RecQ family of DNA helicases. Since 1988, the International Registry of Werner syndrome had enrolled 130 molecularly confirmed WS cases from among 110 worldwide pedigrees. We now report 18 new mutations, including two genomic rearrangements, a deep intronic mutation resulting in a novel exon, a splice consensus mutation leading to utilization of the nearby splice site, and two rare missense mutations. We also review evidence for founder mutations among various ethnic/geographic groups. Founder WRN mutations had been previously reported in Japan and Northern Sardinia. Our Registry now suggests characteristic mutations originated in Morocco, Turkey, The Netherlands and elsewhere.
Figures
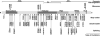
Similar articles
-
The clinical characteristics of Werner syndrome: molecular and biochemical diagnosis.Hum Genet. 2008 Nov;124(4):369-77. doi: 10.1007/s00439-008-0562-0. Epub 2008 Sep 23. Hum Genet. 2008. PMID: 18810497 Free PMC article. Review.
-
Homozygous and compound heterozygous mutations at the Werner syndrome locus.Hum Mol Genet. 1996 Dec;5(12):1909-13. doi: 10.1093/hmg/5.12.1909. Hum Mol Genet. 1996. PMID: 8968742
-
Werner syndrome and mutations of the WRN and LMNA genes in France.Hum Mutat. 2006 Jul;27(7):718-9. doi: 10.1002/humu.9435. Hum Mutat. 2006. PMID: 16786514
-
Mutations in the consensus helicase domains of the Werner syndrome gene. Werner's Syndrome Collaborative Group.Am J Hum Genet. 1997 Feb;60(2):330-41. Am J Hum Genet. 1997. PMID: 9012406 Free PMC article.
-
Search and insights into novel genetic alterations leading to classical and atypical Werner syndrome.Gerontology. 2014;60(3):239-46. doi: 10.1159/000356030. Epub 2014 Jan 3. Gerontology. 2014. PMID: 24401204 Free PMC article. Review.
Cited by
-
Analysis of Pathogenic Pseudoexons Reveals Novel Mechanisms Driving Cryptic Splicing.Front Genet. 2022 Jan 24;12:806946. doi: 10.3389/fgene.2021.806946. eCollection 2021. Front Genet. 2022. PMID: 35140743 Free PMC article.
-
Calcification in Werner syndrome associated with lymphatic vessels aging.Aging (Albany NY). 2021 Dec 27;13(24):25717-25728. doi: 10.18632/aging.203789. Epub 2021 Dec 27. Aging (Albany NY). 2021. PMID: 34958633 Free PMC article.
-
Role of pseudoexons and pseudointrons in human cancer.Int J Cell Biol. 2013;2013:810572. doi: 10.1155/2013/810572. Epub 2013 Sep 24. Int J Cell Biol. 2013. PMID: 24204383 Free PMC article. Review.
-
Functional deficit associated with a missense Werner syndrome mutation.DNA Repair (Amst). 2013 Jun 1;12(6):414-21. doi: 10.1016/j.dnarep.2013.03.004. Epub 2013 Apr 11. DNA Repair (Amst). 2013. PMID: 23583337 Free PMC article.
-
Disease-causing missense mutations in human DNA helicase disorders.Mutat Res. 2013 Apr-Jun;752(2):138-152. doi: 10.1016/j.mrrev.2012.12.004. Epub 2012 Dec 28. Mutat Res. 2013. PMID: 23276657 Free PMC article. Review.
References
-
- Brosh RM, Jr, Opresko PL, Bohr VA. Enzymatic mechanism of the WRN helicase/nuclease. Methods Enzymol. 2006;409:52–85. - PubMed
-
- Chen L, Lee L, Kudlow BA, Dos Santos HG, Sletvold O, Shafeghati Y, Botha EG, Garg A, Hanson NB, Martin GM, Mian IS, Kennedy BK, Oshima J. LMNA mutations in atypical Werner's syndrome. Lancet. 2003;362:440–445. - PubMed
-
- Epstein CJ, Martin GM, Schultz AL, Motulsky AG. Werner's syndrome a review of its symptomatology, natural history, pathologic features, genetics and relationship to the natural aging process. Medicine (Baltimore) 1966;45:177–221. - PubMed
-
- Goto M. Hierarchical deterioration of body systems in Werner's syndrome: implications for normal ageing. Mech Ageing Dev. 1997;98:239–254. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

