Evidence for DNA hairpin recognition by Zta at the Epstein-Barr virus origin of lytic replication
- PMID: 20444899
- PMCID: PMC2898250
- DOI: 10.1128/JVI.02666-09
Evidence for DNA hairpin recognition by Zta at the Epstein-Barr virus origin of lytic replication
Abstract
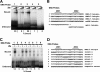
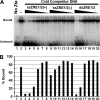
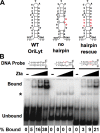
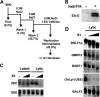
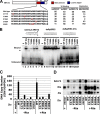
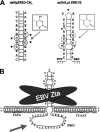
The Epstein-Barr virus immediate-early protein (Zta) plays an essential role in viral lytic activation and pathogenesis. Zta is a basic zipper (b-Zip) domain-containing protein that binds multiple sites in the viral origin of lytic replication (OriLyt) and is required for lytic-cycle DNA replication. We present evidence that Zta binds to a sequence-specific, imperfect DNA hairpin formed by an inverted repeat within the upstream essential element (UEE) of OriLyt. Mutations in the OriLyt sequence that are predicted to disrupt hairpin formation also disrupt Zta binding in vitro. Restoration of the hairpin rescues the defect. We also show that OriLyt DNA isolated from replicating cells contains a nuclease-sensitive region that overlaps with the inverted-repeat region of the UEE. Furthermore, point mutations in Zta that disrupt specific recognition of the UEE hairpin are defective for activation of lytic replication. These data suggest that Zta acts by inducing and/or stabilizing a DNA hairpin structure during productive infection. The DNA hairpin at OriLyt with which Zta interacts resembles DNA structures formed at other herpesvirus origins and may therefore represent a common secondary structure used by all herpesvirus family members during the initiation of DNA replication.
Figures

References
-
- Andersson, J. 2000. An overview of Epstein-Barr virus: from discovery to future directions for treatment and prevention. Herpes 7:76-82. - PubMed
-
- Arbuckle, M. I., and N. D. Stow. 1993. A mutational analysis of the DNA-binding domain of the herpes simplex virus type 1 UL9 protein. J. Gen. Virol. 74(Pt. 7):1349-1355. - PubMed
-
- Aslani, A., M. Olsson, and P. Elias. 2002. ATP-dependent unwinding of a minimal origin of DNA replication by the origin-binding protein and the single-strand DNA-binding protein ICP8 from herpes simplex virus type I. J. Biol. Chem. 277:41204-41212. - PubMed
-
- Aslani, A., S. Simonsson, and P. Elias. 2000. A novel conformation of the herpes simplex virus origin of DNA replication recognized by the origin binding protein. J. Biol. Chem. 275:5880-5887. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

