The ubiquitin/26S proteasome system in plant-pathogen interactions: a never-ending hide-and-seek game
- PMID: 20447278
- PMCID: PMC6640532
- DOI: 10.1111/j.1364-3703.2009.00596.x
The ubiquitin/26S proteasome system in plant-pathogen interactions: a never-ending hide-and-seek game
Erratum in
- Mol Plant Pathol. 2011 Jan;12(1):103
Abstract
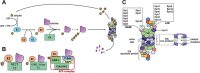
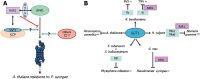
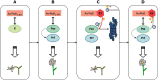
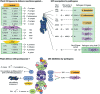
The ubiquitin/26S proteasome system (UPS) plays a central role in plant protein degradation. Over the past few years, the importance of this pathway in plant-pathogen interactions has been increasingly highlighted. UPS is involved in almost every step of the defence mechanisms in plants, regardless of the type of pathogen. In addition to its proteolytic activities, UPS, through its 20S RNase activity, may be part of a still unknown antiviral defence pathway. Strikingly, UPS is not only a weapon used by plants to defend themselves, but also a target for some pathogens that have evolved mechanisms to inhibit and/or use this system for their own purposes. This article attempts to summarize the current knowledge on UPS involvement in plant-microbe interactions, a complex scheme that illustrates the never-ending arms race between hosts and microbes.
Figures
Similar articles
-
Plant Virus Infection and the Ubiquitin Proteasome Machinery: Arms Race along the Endoplasmic Reticulum.Viruses. 2016 Nov 19;8(11):314. doi: 10.3390/v8110314. Viruses. 2016. PMID: 27869775 Free PMC article. Review.
-
Manipulation of host proteasomes as a virulence mechanism of plant pathogens.Annu Rev Phytopathol. 2013;51:521-42. doi: 10.1146/annurev-phyto-082712-102312. Epub 2013 May 31. Annu Rev Phytopathol. 2013. PMID: 23725468 Review.
-
The ubiquitin-proteasome system in plant responses to environments.Plant Cell Environ. 2019 Oct;42(10):2931-2944. doi: 10.1111/pce.13633. Epub 2019 Aug 13. Plant Cell Environ. 2019. PMID: 31364170 Review.
-
The ubiquitin-proteasome system in positive-strand RNA virus infection.Rev Med Virol. 2013 Mar;23(2):85-96. doi: 10.1002/rmv.1725. Epub 2012 Jul 11. Rev Med Virol. 2013. PMID: 22782620 Free PMC article. Review.
-
Regulation of Plant Immunity by the Proteasome.Int Rev Cell Mol Biol. 2019;343:37-63. doi: 10.1016/bs.ircmb.2018.06.004. Epub 2018 Aug 1. Int Rev Cell Mol Biol. 2019. PMID: 30712674 Review.
Cited by
-
iTRAQ-based comparative proteome analyses of different growth stages revealing the regulatory role of reactive oxygen species in the fruiting body development of Ophiocordyceps sinensis.PeerJ. 2021 Mar 3;9:e10940. doi: 10.7717/peerj.10940. eCollection 2021. PeerJ. 2021. PMID: 33717691 Free PMC article.
-
A viral deubiquitylating enzyme targets viral RNA-dependent RNA polymerase and affects viral infectivity.EMBO J. 2012 Feb 1;31(3):741-53. doi: 10.1038/emboj.2011.424. Epub 2011 Nov 25. EMBO J. 2012. PMID: 22117220 Free PMC article.
-
Identification of the interacting proteins of Bambusa pervariabilis × Dendrocalamopsis grandis in response to the transcription factor ApCtf1β in Arthrinium phaeospermum.Front Plant Sci. 2022 Sep 15;13:991077. doi: 10.3389/fpls.2022.991077. eCollection 2022. Front Plant Sci. 2022. PMID: 36186076 Free PMC article.
-
Genome-wide identification, phylogenetic and expression analysis of the maize HECT E3 ubiquitin ligase genes.Genetica. 2019 Dec;147(5-6):391-400. doi: 10.1007/s10709-019-00080-4. Epub 2019 Nov 18. Genetica. 2019. PMID: 31741104
-
The E3 Ubiquitin Ligase ATL9 Affects Expression of Defense Related Genes, Cell Death and Callose Deposition in Response to Fungal Infection.Pathogens. 2022 Jan 5;11(1):68. doi: 10.3390/pathogens11010068. Pathogens. 2022. PMID: 35056016 Free PMC article.
References
-
- Angot, A. , Peeters, N. , Lechner, E. , Vailleau, F. , Baud, C. , Gentzbittel, L. , Sartorel, E. , Genschik, P. , Boucher, C. and Genin, S. (2006) Ralstonia solanacearum requires F‐box‐like domain‐containing type III effectors to promote disease on several host plants. PNAS, 103, 14620–14625. - PMC - PubMed
-
- Apcher, G.S. , Heink, S. , Zantopf, D. , Kloetzel, P.‐M. , Schmid, H.‐P. , Mayer, R.J. and Kruger, E. (2003) Human immunodeficiency virus‐1 Tat protein interacts with distinct proteasomal [alpha] and [beta] subunits. FEBS Letters, 553, 200–204. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

