Metallo-aminopeptidase inhibitors
- PMID: 20457213
- PMCID: PMC7117057
- DOI: 10.1016/j.biochi.2010.04.026
Metallo-aminopeptidase inhibitors
Abstract
Aminopeptidases are enzymes that selectively hydrolyze an amino acid residue from the N-terminus of proteins and peptides. They are important for the proper functioning of prokaryotic and eukaryotic cells, but very often are central players in the devastating human diseases like cancer, malaria and diabetes. The largest aminopeptidase group include enzymes containing metal ion(s) in their active centers, which often determines the type of inhibitors that are the most suitable for them. Effective ligands mostly bind in a non-covalent mode by forming complexes with the metal ion(s). Here, we present several approaches for the design of inhibitors for metallo-aminopeptidases. The optimized structures should be considered as potential leads in the drug discovery process against endogenous and infectious diseases.
Crown Copyright © 2010. Published by Elsevier Masson SAS. All rights reserved.
Figures
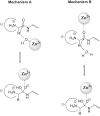
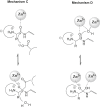
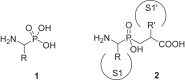
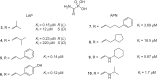

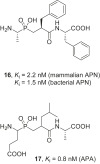




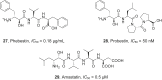


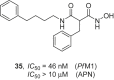
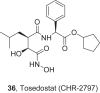


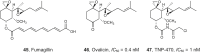


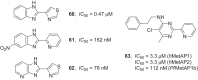
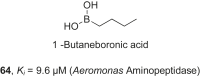

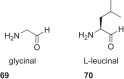
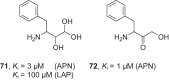

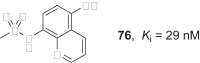
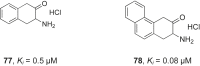

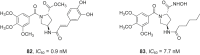



References
-
- Hooper N.M., Lendeckel U. vol. 2. Kluwer Academic/Plenum Publishers; New York: 2004. (Aminopeptidases in Biology and Disease, Proteases in Biology and Disease).
-
- Sanderink G.J., Artur Y., Siest G. Human aminopeptidases: a review of the literature. J. Clin. Chem. Clin. Biochem. 1988;26:795–807. - PubMed
-
- Sanz Y. Aminopeptidases. In: Polaina J., MacCabe A.P., editors. Industrial Enzymes. Springer; 2007. pp. 243–260.
-
- Jankiewicz U., Bielawski W. The properties and functions of bacterial aminopeptidases. Acta Microbiol. Pol. 2003;52:217–231. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

