Hepatitis C virus: assembly and release of virus particles
- PMID: 20457608
- PMCID: PMC2906262
- DOI: 10.1074/jbc.R110.133017
Hepatitis C virus: assembly and release of virus particles
Abstract
Hepatitis C virus is a blood-borne virus that typically establishes a chronic infection in the liver, which often results in cirrhosis and hepatocellular carcinoma. Progress in understanding the complete virus life cycle has been greatly enhanced by the recent availability of a tissue culture system that produces infectious virus progeny. Thus, it is now possible to gain insight into the roles played by viral components in assembly and egress and the cellular pathways that contribute to virion formation. This minireview describes the key determining viral and host factors that are needed to produce infectious virus.
Figures


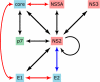
Similar articles
-
Cooperation between the Hepatitis C Virus p7 and NS5B Proteins Enhances Virion Infectivity.J Virol. 2015 Nov;89(22):11523-33. doi: 10.1128/JVI.01185-15. Epub 2015 Sep 9. J Virol. 2015. PMID: 26355084 Free PMC article.
-
Roles of Lipoproteins and Apolipoproteins in Particle Formation of Hepatitis C Virus.Trends Microbiol. 2015 Oct;23(10):618-629. doi: 10.1016/j.tim.2015.07.007. Trends Microbiol. 2015. PMID: 26433694 Review.
-
Host-derived apolipoproteins play comparable roles with viral secretory proteins Erns and NS1 in the infectious particle formation of Flaviviridae.PLoS Pathog. 2017 Jun 23;13(6):e1006475. doi: 10.1371/journal.ppat.1006475. eCollection 2017 Jun. PLoS Pathog. 2017. PMID: 28644867 Free PMC article.
-
Visualizing the Essential Role of Complete Virion Assembly Machinery in Efficient Hepatitis C Virus Cell-to-Cell Transmission by a Viral Infection-Activated Split-Intein-Mediated Reporter System.J Virol. 2017 Jan 3;91(2):e01720-16. doi: 10.1128/JVI.01720-16. Print 2017 Jan 15. J Virol. 2017. PMID: 27852847 Free PMC article.
-
Virion assembly and release.Curr Top Microbiol Immunol. 2013;369:199-218. doi: 10.1007/978-3-642-27340-7_8. Curr Top Microbiol Immunol. 2013. PMID: 23463202 Free PMC article. Review.
Cited by
-
Lipid Droplets Accumulation during Hepatitis C Virus Infection in Cell-Culture Varies among Genotype 1-3 Strains and Does Not Correlate with Virus Replication.Viruses. 2021 Feb 28;13(3):389. doi: 10.3390/v13030389. Viruses. 2021. PMID: 33671086 Free PMC article.
-
Lipoprotein lipase inhibits hepatitis C virus (HCV) infection by blocking virus cell entry.PLoS One. 2011;6(10):e26637. doi: 10.1371/journal.pone.0026637. Epub 2011 Oct 21. PLoS One. 2011. PMID: 22039521 Free PMC article.
-
Extracellular Interactions between Hepatitis C Virus and Secreted Apolipoprotein E.J Virol. 2017 Jul 12;91(15):e02227-16. doi: 10.1128/JVI.02227-16. Print 2017 Aug 1. J Virol. 2017. PMID: 28539442 Free PMC article.
-
Hepatitis C Virus-Lipid Interplay: Pathogenesis and Clinical Impact.Biomedicines. 2023 Jan 19;11(2):271. doi: 10.3390/biomedicines11020271. Biomedicines. 2023. PMID: 36830808 Free PMC article. Review.
-
Packaging of Genomic RNA in Positive-Sense Single-Stranded RNA Viruses: A Complex Story.Viruses. 2019 Mar 13;11(3):253. doi: 10.3390/v11030253. Viruses. 2019. PMID: 30871184 Free PMC article. Review.
References
-
- Kim Y. K., Kim C. S., Lee S. H., Jang S. K. (2002) Biochem. Biophys. Res. Commun. 290, 105–112 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

