The archaeal transamidosome for RNA-dependent glutamine biosynthesis
- PMID: 20457752
- PMCID: PMC2943598
- DOI: 10.1093/nar/gkq336
The archaeal transamidosome for RNA-dependent glutamine biosynthesis
Abstract
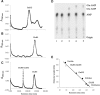
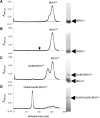
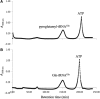
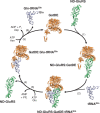
Archaea make glutaminyl-tRNA (Gln-tRNA(Gln)) in a two-step process; a non-discriminating glutamyl-tRNA synthetase (ND-GluRS) forms Glu-tRNA(Gln), while the heterodimeric amidotransferase GatDE converts this mischarged tRNA to Gln-tRNA(Gln). Many prokaryotes synthesize asparaginyl-tRNA (Asn-tRNA(Asn)) in a similar manner using a non-discriminating aspartyl-tRNA synthetase (ND-AspRS) and the heterotrimeric amidotransferase GatCAB. The transamidosome, a complex of tRNA synthetase, amidotransferase and tRNA, was first described for the latter system in Thermus thermophilus [Bailly, M., Blaise, M., Lorber, B., Becker, H.D. and Kern, D. (2007) The transamidosome: a dynamic ribonucleoprotein particle dedicated to prokaryotic tRNA-dependent asparagine biosynthesis. Mol. Cell, 28, 228-239.]. Here, we show a similar complex for Gln-tRNA(Gln) formation in Methanothermobacter thermautotrophicus that allows the mischarged Glu-tRNA(Gln) made by the tRNA synthetase to be channeled to the amidotransferase. The association of archaeal ND-GluRS with GatDE (K(D) = 100 ± 22 nM) sequesters the tRNA synthetase for Gln-tRNA(Gln) formation, with GatDE reducing the affinity of ND-GluRS for tRNA(Glu) by at least 13-fold. Unlike the T. thermophilus transamidosome, the archaeal complex does not require tRNA for its formation, is not stable through product (Gln-tRNA(Gln)) formation, and has no major effect on the kinetics of tRNA(Gln) glutamylation nor transamidation. The differences between the two transamidosomes may be a consequence of the fact that ND-GluRS is a class I aminoacyl-tRNA synthetase, while ND-AspRS belongs to the class II family.
Figures


Similar articles
-
Gln-tRNAGln synthesis in a dynamic transamidosome from Helicobacter pylori, where GluRS2 hydrolyzes excess Glu-tRNAGln.Nucleic Acids Res. 2011 Nov;39(21):9306-15. doi: 10.1093/nar/gkr619. Epub 2011 Aug 3. Nucleic Acids Res. 2011. PMID: 21813455 Free PMC article.
-
Two enzymes bound to one transfer RNA assume alternative conformations for consecutive reactions.Nature. 2010 Sep 30;467(7315):612-6. doi: 10.1038/nature09411. Nature. 2010. PMID: 20882017
-
Methanothermobacter thermautotrophicus tRNA Gln confines the amidotransferase GatCAB to asparaginyl-tRNA Asn formation.J Mol Biol. 2008 Mar 28;377(3):845-53. doi: 10.1016/j.jmb.2008.01.064. Epub 2008 Jan 31. J Mol Biol. 2008. PMID: 18291416 Free PMC article.
-
Divergence of glutamate and glutamine aminoacylation pathways: providing the evolutionary rationale for mischarging.J Mol Evol. 1995 May;40(5):476-81. doi: 10.1007/BF00166615. J Mol Evol. 1995. PMID: 7783222 Review.
-
Glutamyl-tRNA sythetase.Biol Chem. 1997 Nov;378(11):1313-29. Biol Chem. 1997. PMID: 9426192 Review.
Cited by
-
Synthesis of Glu-tRNA(Gln) by engineered and natural aminoacyl-tRNA synthetases.Biochemistry. 2010 Aug 10;49(31):6727-36. doi: 10.1021/bi100886z. Biochemistry. 2010. PMID: 20617848 Free PMC article.
-
Evolution of Life on Earth: tRNA, Aminoacyl-tRNA Synthetases and the Genetic Code.Life (Basel). 2020 Mar 2;10(3):21. doi: 10.3390/life10030021. Life (Basel). 2020. PMID: 32131473 Free PMC article. Review.
-
Gln-tRNAGln synthesis in a dynamic transamidosome from Helicobacter pylori, where GluRS2 hydrolyzes excess Glu-tRNAGln.Nucleic Acids Res. 2011 Nov;39(21):9306-15. doi: 10.1093/nar/gkr619. Epub 2011 Aug 3. Nucleic Acids Res. 2011. PMID: 21813455 Free PMC article.
-
Non-canonical roles of tRNAs and tRNA mimics in bacterial cell biology.Mol Microbiol. 2016 Aug;101(4):545-58. doi: 10.1111/mmi.13419. Epub 2016 Jun 28. Mol Microbiol. 2016. PMID: 27169680 Free PMC article. Review.
-
Two enzymes bound to one transfer RNA assume alternative conformations for consecutive reactions.Nature. 2010 Sep 30;467(7315):612-6. doi: 10.1038/nature09411. Nature. 2010. PMID: 20882017
References
-
- Ibba M, Söll D. Aminoacyl-tRNA synthesis. Annu. Rev. Biochem. 2000;69:617–650. - PubMed
-
- Becker HD, Reinbolt J, Kreutzer R, Giegé R, Kern D. Existence of two distinct aspartyl-tRNA synthetases in Thermus thermophilus. Structural and biochemical properties of the two enzymes. Biochemistry. 1997;36:8785–8797. - PubMed

