Spatio-temporal modeling of signaling protein recruitment to EGFR
- PMID: 20459599
- PMCID: PMC2877007
- DOI: 10.1186/1752-0509-4-57
Spatio-temporal modeling of signaling protein recruitment to EGFR
Abstract
Background: A stochastic simulator was implemented to study EGFR signal initiation in 3D with single molecule detail. The model considers previously unexplored contributions to receptor-adaptor coupling, such as receptor clustering and diffusive properties of both receptors and binding partners. The agent-based and rule-based approach permits consideration of combinatorial complexity, a problem associated with multiple phosphorylation sites and the potential for simultaneous binding of adaptors.
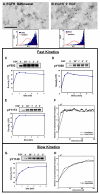
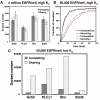
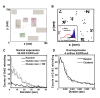
Results: The model was used to simulate recruitment of four different signaling molecules (Grb2, PLCgamma1, Stat5, Shc) to the phosphorylated EGFR tail, with rules based on coarse-grained prediction of spatial constraints. Parameters were derived in part from quantitative immunoblotting, immunoprecipitation and electron microscopy data. Results demonstrate that receptor clustering increases the efficiency of individual adaptor retainment on activated EGFR, an effect that is overridden if crowding is imposed by receptor overexpression. Simultaneous docking of multiple proteins is highly dependent on receptor-adaptor stability and independent of clustering.
Conclusions: Overall, we propose that receptor density, reaction kinetics and membrane spatial organization all contribute to signaling efficiency and influence the carcinogenesis process.
Figures


References
-
- Ushiro H, Cohen S. Identification of phosphotyrosine as a product of epidermal growth factor-activated protein kinase in A-431 cell membranes. J Biol Chem. 1980;255:8363–8365. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

