Modeling non-uniformity in short-read rates in RNA-Seq data
- PMID: 20459815
- PMCID: PMC2898062
- DOI: 10.1186/gb-2010-11-5-r50
Modeling non-uniformity in short-read rates in RNA-Seq data
Abstract
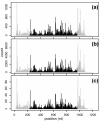
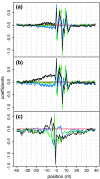
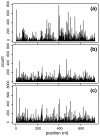
After mapping, RNA-Seq data can be summarized by a sequence of read counts commonly modeled as Poisson variables with constant rates along each transcript, which actually fit data poorly. We suggest using variable rates for different positions, and propose two models to predict these rates based on local sequences. These models explain more than 50% of the variations and can lead to improved estimates of gene and isoform expressions for both Illumina and Applied Biosystems data.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

