The BMP pathway acts to directly regulate Tbx20 in the developing heart
- PMID: 20460370
- PMCID: PMC2867324
- DOI: 10.1242/dev.043588
The BMP pathway acts to directly regulate Tbx20 in the developing heart
Abstract
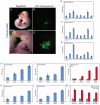
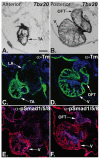
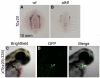
TBX20 has been shown to be essential for vertebrate heart development. Mutations within the TBX20 coding region are associated with human congenital heart disease, and the loss of Tbx20 in a wide variety of model systems leads to cardiac defects and eventually heart failure. Despite the crucial role of TBX20 in a range of cardiac cellular processes, the signal transduction pathways that act upstream of Tbx20 remain unknown. Here, we have identified and characterized a conserved 334 bp Tbx20 cardiac regulatory element that is directly activated by the BMP/SMAD1 signaling pathway. We demonstrate that this element is both necessary and sufficient to drive cardiac-specific expression of Tbx20 in Xenopus, and that blocking SMAD1 signaling in vivo specifically abolishes transcription of Tbx20, but not that of other cardiac factors, such as Tbx5 and MHC, in the developing heart. We further demonstrate that activation of Tbx20 by SMAD1 is mediated by a set of novel, non-canonical, high-affinity SMAD-binding sites located within this regulatory element and that phospho-SMAD1 directly binds a non-canonical SMAD1 site in vivo. Finally, we show that these non-canonical sites are necessary and sufficient for Tbx20 expression in Xenopus, and that reporter constructs containing these sites are expressed in a cardiac-specific manner in zebrafish and mouse. Collectively, our findings define Tbx20 as a direct transcriptional target of the BMP/SMAD1 signaling pathway during cardiac maturation.
Figures





References
-
- Ahn D. G., Ruvinsky I., Oates A. C., Silver L. M., Ho R. K. (2000). tbx20, a new vertebrate T-box gene expressed in the cranial motor neurons and developing cardiovascular structures in zebrafish. Mech. Dev. 95, 253-258 - PubMed
-
- Baldini A. (2004). DiGeorge syndrome: an update. Curr. Opin. Cardiol. 19, 201-204 - PubMed
-
- Bauer H., Lele Z., Rauch G. J., Geisler R., Hammerschmidt M. (2001). The type I serine/threonine kinase receptor Alk8/Lost-a-fin is required for Bmp2b/7 signal transduction during dorsoventral patterning of the zebrafish embryo. Development 128, 849-858 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

