Structure and mechanism of receptor sharing by the IL-10R2 common chain
- PMID: 20462497
- PMCID: PMC2879597
- DOI: 10.1016/j.str.2010.02.009
Structure and mechanism of receptor sharing by the IL-10R2 common chain
Abstract
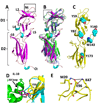
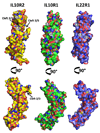
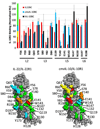
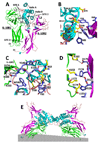
IL-10R2 is a shared cell surface receptor required for the activation of five class 2 cytokines (IL-10, IL-22, IL-26, IL-28, and IL-29) that play critical roles in host defense. To define the molecular mechanisms that regulate its promiscuous binding, we have determined the crystal structure of the IL-10R2 ectodomain at 2.14 A resolution. IL-10R2 residues required for binding were identified by alanine scanning and used to derive computational models of IL-10/IL-10R1/IL-10R2 and IL-22/IL-22R1/IL-10R2 ternary complexes. The models reveal a conserved binding epitope that is surrounded by two clefts that accommodate the structural and chemical diversity of the cytokines. These results provide a structural framework for interpreting IL-10R2 single nucleotide polymorphisms associated with human disease.
Copyright 2010 Elsevier Ltd. All rights reserved.
Figures


Comment in
-
Class II cytokine common receptors: something old, something new.Structure. 2010 May 12;18(5):551-2. doi: 10.1016/j.str.2010.04.003. Structure. 2010. PMID: 20462488
Similar articles
-
Conformational changes mediate interleukin-10 receptor 2 (IL-10R2) binding to IL-10 and assembly of the signaling complex.J Biol Chem. 2006 Nov 17;281(46):35088-96. doi: 10.1074/jbc.M606791200. Epub 2006 Sep 18. J Biol Chem. 2006. PMID: 16982608
-
A model of the ternary complex of interleukin-10 with its soluble receptors.BMC Struct Biol. 2005 Jun 28;5:10. doi: 10.1186/1472-6807-5-10. BMC Struct Biol. 2005. PMID: 15985167 Free PMC article.
-
Is there an interaction between interleukin-10 and interleukin-22?Genes Immun. 2005 Feb;6(1):8-18. doi: 10.1038/sj.gene.6364144. Genes Immun. 2005. PMID: 15526001
-
The expanded family of class II cytokines that share the IL-10 receptor-2 (IL-10R2) chain.J Leukoc Biol. 2004 Aug;76(2):314-21. doi: 10.1189/jlb.0204117. Epub 2004 May 3. J Leukoc Biol. 2004. PMID: 15123776 Review.
-
Structural features of the interleukin-10 family of cytokines.Curr Pharm Des. 2004;10(31):3873-84. doi: 10.2174/1381612043382602. Curr Pharm Des. 2004. PMID: 15579076 Review.
Cited by
-
Crosstalk of Inflammatory Cytokines within the Breast Tumor Microenvironment.Int J Mol Sci. 2023 Feb 16;24(4):4002. doi: 10.3390/ijms24044002. Int J Mol Sci. 2023. PMID: 36835413 Free PMC article. Review.
-
A variant upstream of IFNL3 (IL28B) creating a new interferon gene IFNL4 is associated with impaired clearance of hepatitis C virus.Nat Genet. 2013 Feb;45(2):164-71. doi: 10.1038/ng.2521. Epub 2013 Jan 6. Nat Genet. 2013. PMID: 23291588 Free PMC article.
-
Synthetic interleukin 22 (IL-22) signaling reveals biological activity of homodimeric IL-10 receptor 2 and functional cross-talk with the IL-6 receptor gp130.J Biol Chem. 2020 Aug 28;295(35):12378-12397. doi: 10.1074/jbc.RA120.013927. Epub 2020 Jul 1. J Biol Chem. 2020. PMID: 32611765 Free PMC article.
-
Molecular Mechanics Simulations and Experimental Investigation of the Effect of Tadalafil on Various Inflammatory Pain Mediators.ACS Omega. 2022 Nov 22;7(48):43747-43758. doi: 10.1021/acsomega.2c04761. eCollection 2022 Dec 6. ACS Omega. 2022. PMID: 36506154 Free PMC article.
-
Interleukin-10 paradox: A potent immunoregulatory cytokine that has been difficult to harness for immunotherapy.Cytokine. 2015 Jul;74(1):27-34. doi: 10.1016/j.cyto.2014.10.031. Epub 2014 Dec 4. Cytokine. 2015. PMID: 25481648 Free PMC article. Review.
References
-
- Abrahams JP, Leslie AG. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr D Biol Crystallogr. 1996;52:30–42. - PubMed
-
- Blackburn SD, Wherry EJ. IL-10, T cell exhaustion and viral persistence. Trends Microbiol. 2007;15:143–146. - PubMed
-
- Bleicher L, de Moura PR, Watanabe L, Colau D, Dumoutier L, Renauld JC, Polikarpov I. Crystal structure of the IL-22/IL-22R1 complex and its implications for the IL-22 signaling mechanism. FEBS Lett. 2008;582:2985–2992. - PubMed
-
- Boulanger MJ, Chow DC, Brevnova EE, Garcia KC. Hexameric structure and assembly of the interleukin-6/IL-6 alpha-receptor/gp130 complex. Science. 2003;300:2101–2104. - PubMed
-
- Brown RJ, Adams JJ, Pelekanos RA, Wan Y, McKinstry WJ, Palethorpe K, Seeber RM, Monks TA, Eidne KA, Parker MW, Waters MJ. Model for growth hormone receptor activation based on subunit rotation within a receptor dimer. Nat Struct Mol Biol. 2005;12:814–821. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

