APOBEC3G contributes to HIV-1 variation through sublethal mutagenesis
- PMID: 20463080
- PMCID: PMC2898230
- DOI: 10.1128/JVI.00056-10
APOBEC3G contributes to HIV-1 variation through sublethal mutagenesis
Abstract
The mammalian APOBEC3 proteins are an important component of the cellular innate immune response to retroviral infection. APOBEC3G can extinguish HIV-1 infectivity by its incorporation into virus particles and subsequent cytosine deaminase activity that attacks the nascent viral cDNA during reverse transcription, causing lethal mutagenesis. It has been suggested, but not formally shown, that APOBEC3G can also induce sublethal mutagenesis, which would maintain virus infectivity and contribute to HIV-1 variation. To test this, we developed a novel model system utilizing an HIV-1 vector and a panel of APOBEC3G-expressing cells. We observed proviruses with single APOBEC3G-mediated mutations (in the presence or absence of Vif), occurring at distinct hot spots and which could be rescued and shown to have infectivity. These data indicate that APOBEC3G-dependent restriction of HIV-1 can result in viable viral progeny that harbor sublethal levels of G-to-A mutations. Such mutations have the potential to contribute significantly to HIV-1 evolution, pathogenesis, immune escape, and drug resistance.
Figures
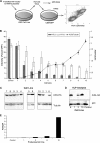
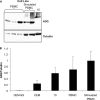
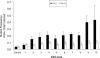
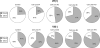

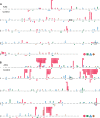
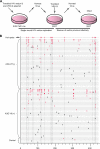
References
-
- Bebenek, K., J. Abbotts, J. D. Roberts, S. H. Wilson, and T. A. Kunkel. 1989. Specificity and mechanism of error-prone replication by human immunodeficiency virus-1 reverse transcriptase. J. Biol. Chem. 264:16948-16956. - PubMed
-
- Bebenek, K., J. D. Roberts, and T. A. Kunkel. 1992. The effects of dNTP pool imbalances on frameshift fidelity during DNA replication. J. Biol. Chem. 267:3589-3596. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

