Degradation of YRA1 Pre-mRNA in the cytoplasm requires translational repression, multiple modular intronic elements, Edc3p, and Mex67p
- PMID: 20463951
- PMCID: PMC2864733
- DOI: 10.1371/journal.pbio.1000360
Degradation of YRA1 Pre-mRNA in the cytoplasm requires translational repression, multiple modular intronic elements, Edc3p, and Mex67p
Erratum in
-
Correction: Degradation of YRA1 Pre-mRNA in the Cytoplasm Requires Translational Repression, Multiple Modular Intronic Elements, Edc3p, and Mex67p.PLoS Biol. 2016 May 12;14(5):e1002470. doi: 10.1371/journal.pbio.1002470. eCollection 2016 May. PLoS Biol. 2016. PMID: 27171161 Free PMC article.
Abstract
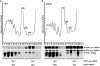
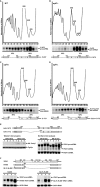
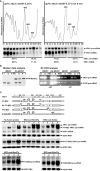
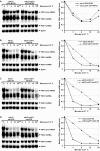
Intron-containing pre-mRNAs are normally retained and processed in the nucleus but are sometimes exported to the cytoplasm and degraded by the nonsense-mediated mRNA decay (NMD) pathway as a consequence of their inclusion of intronic in-frame termination codons. When shunted to the cytoplasm by autoregulated nuclear export, the intron-containing yeast YRA1 pre-mRNA evades NMD and is targeted by a cytoplasmic decay pathway mediated by the decapping activator Edc3p. Here, we have elucidated this transcript-specific decay mechanism, showing that Edc3p-mediated YRA1 pre-mRNA degradation occurs independently of translation and is controlled through five structurally distinct but functionally interdependent modular elements in the YRA1 intron. Two of these elements target the pre-mRNA as an Edc3p substrate and the other three mediate transcript-specific translational repression. Translational repression of YRA1 pre-mRNA also requires the heterodimeric Mex67p/Mtr2p general mRNA export receptor, but not Edc3p, and serves to enhance Edc3p substrate specificity by inhibiting the susceptibility of this pre-mRNA to NMD. Collectively, our data indicate that YRA1 pre-mRNA degradation is a highly regulated process that proceeds through translational repression, substrate recognition by Edc3p, recruitment of the Dcp1p/Dcp2p decapping enzyme, and activation of decapping.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Similar articles
-
YRA1 autoregulation requires nuclear export and cytoplasmic Edc3p-mediated degradation of its pre-mRNA.Mol Cell. 2007 Feb 23;25(4):559-73. doi: 10.1016/j.molcel.2007.01.012. Mol Cell. 2007. PMID: 17317628 Free PMC article.
-
Expression of the essential mRNA export factor Yra1p is autoregulated by a splicing-dependent mechanism.RNA. 2002 Aug;8(8):969-80. doi: 10.1017/s1355838202020046. RNA. 2002. PMID: 12212852 Free PMC article.
-
Identification of Edc3p as an enhancer of mRNA decapping in Saccharomyces cerevisiae.Genetics. 2004 Feb;166(2):729-39. doi: 10.1534/genetics.166.2.729. Genetics. 2004. PMID: 15020463 Free PMC article.
-
Regulation of mRNA decay: decapping goes solo.Mol Cell. 2004 Jul 2;15(1):1-2. doi: 10.1016/j.molcel.2004.06.031. Mol Cell. 2004. PMID: 15225542 Review.
-
The functional complexity of the RNA-binding protein Yra1: mRNA biogenesis, genome stability and DSB repair.Curr Genet. 2020 Feb;66(1):63-71. doi: 10.1007/s00294-019-01011-8. Epub 2019 Jul 10. Curr Genet. 2020. PMID: 31292684 Review.
Cited by
-
Intra- and intermolecular regulatory interactions in Upf1, the RNA helicase central to nonsense-mediated mRNA decay in yeast.Mol Cell Biol. 2013 Dec;33(23):4672-84. doi: 10.1128/MCB.01136-13. Epub 2013 Oct 7. Mol Cell Biol. 2013. PMID: 24100012 Free PMC article.
-
SUS1 introns are required for efficient mRNA nuclear export in yeast.Nucleic Acids Res. 2011 Oct;39(19):8599-611. doi: 10.1093/nar/gkr496. Epub 2011 Jul 12. Nucleic Acids Res. 2011. PMID: 21749979 Free PMC article.
-
Dcp2 C-terminal cis-binding elements control selective targeting of the decapping enzyme by forming distinct decapping complexes.Elife. 2022 May 23;11:e74410. doi: 10.7554/eLife.74410. Elife. 2022. PMID: 35604319 Free PMC article.
-
Yeast Edc3 targets RPS28B mRNA for decapping by binding to a 3' untranslated region decay-inducing regulatory element.Mol Cell Biol. 2014 Apr;34(8):1438-51. doi: 10.1128/MCB.01584-13. Epub 2014 Feb 3. Mol Cell Biol. 2014. PMID: 24492965 Free PMC article.
-
Pby1 is a direct partner of the Dcp2 decapping enzyme.Nucleic Acids Res. 2020 Jun 19;48(11):6353-6366. doi: 10.1093/nar/gkaa337. Nucleic Acids Res. 2020. PMID: 32396195 Free PMC article.
References
-
- Coller J, Parker R. Eukaryotic mRNA decapping. Annu Rev Biochem. 2004;73:861–890. - PubMed
-
- Amrani N, Sachs M. S, Jacobson A. Early nonsense: mRNA decay solves a translational problem. Nat Rev Mol Cell Biol. 2006;7:415–425. - PubMed
-
- Frischmeyer P. A, van Hoof A, O'Donnell K, Guerrerio A. L, Parker R, et al. An mRNA surveillance mechanism that eliminates transcripts lacking termination codons. Science. 2002;295:2258–2261. - PubMed
-
- van Hoof A, Frischmeyer P. A, Dietz H. C, Parker R. Exosome-mediated recognition and degradation of mRNAs lacking a termination codon. Science. 2002;295:2262–2264. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

