Aurora B phosphorylates spatially distinct targets to differentially regulate the kinetochore-microtubule interface
- PMID: 20471944
- PMCID: PMC2873218
- DOI: 10.1016/j.molcel.2010.02.034
Aurora B phosphorylates spatially distinct targets to differentially regulate the kinetochore-microtubule interface
Abstract
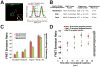
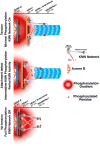
Accurate chromosome segregation requires carefully regulated interactions between kinetochores and microtubules, but how plasticity is achieved to correct diverse attachment defects remains unclear. Here we demonstrate that Aurora B kinase phosphorylates three spatially distinct targets within the conserved outer kinetochore KNL1/Mis12 complex/Ndc80 complex (KMN) network, the key player in kinetochore-microtubule attachments. The combinatorial phosphorylation of the KMN network generates graded levels of microtubule-binding activity, with full phosphorylation severely compromising microtubule binding. Altering the phosphorylation state of each protein causes corresponding chromosome segregation defects. Importantly, the spatial distribution of these targets along the kinetochore axis leads to their differential phosphorylation in response to changes in tension and attachment state. In total, rather than generating exclusively binary changes in microtubule binding, our results suggest a mechanism for the tension-dependent fine-tuning of kinetochore-microtubule interactions.
Copyright (c) 2010 Elsevier Inc. All rights reserved.
Figures




References
-
- Cantin GT, Yates JR., 3rd Strategies for shotgun identification of post-translational modifications by mass spectrometry. J Chromatogr A. 2004;1053:7–14. - PubMed
-
- Cheeseman IM, Chappie JS, Wilson-Kubalek EM, Desai A. The conserved KMN network constitutes the core microtubule-binding site of the kinetochore. Cell. 2006;127:983–997. - PubMed
-
- Cheeseman IM, Desai A. Molecular architecture of the kinetochore-microtubule interface. Nat Rev Mol Cell Biol. 2008;9:33–46. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

