The Saccharomyces cerevisiae RAD9, RAD17 and RAD24 genes are required for suppression of mutagenic post-replicative repair during chronic DNA damage
- PMID: 20472512
- PMCID: PMC2893243
- DOI: 10.1016/j.dnarep.2010.04.007
The Saccharomyces cerevisiae RAD9, RAD17 and RAD24 genes are required for suppression of mutagenic post-replicative repair during chronic DNA damage
Abstract
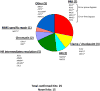
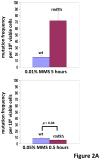
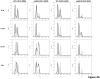
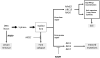
In Saccharomyces cerevisiae, a DNA damage checkpoint in the S-phase is responsible for delaying DNA replication in response to genotoxic stress. This pathway is partially regulated by the checkpoint proteins Rad9, Rad17 and Rad24. Here, we describe a novel hypermutable phenotype for rad9Delta, rad17Delta and rad24Delta cells in response to a chronic 0.01% dose of the DNA alkylating agent MMS. We report that this hypermutability results from DNA damage introduction during the S-phase and is dependent on a functional translesion synthesis pathway. In addition, we performed a genetic screen for interactions with rad9Delta that confer sensitivity to 0.01% MMS. We report and quantify 25 genetic interactions with rad9Delta, many of which involve the post-replication repair machinery. From these data, we conclude that defects in S-phase checkpoint regulation lead to increased reliance on mutagenic translesion synthesis, and we describe a novel role for members of the S-phase DNA damage checkpoint in suppressing mutagenic post-replicative repair in response to sublethal MMS treatment.
2010 Elsevier B.V. All rights reserved.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures




Similar articles
-
[Interaction between checkpoint genes RAD9, RAD17, RAD24, and RAD53 involved in the determination of yeast Saccharomyces cerevisiae sensitivity to ionizing radiation].Genetika. 2008 Jun;44(6):761-70. Genetika. 2008. PMID: 18727386 Russian.
-
[Interaction between checkpoint genes RAD9, RAD17, RAD24, and RAD53 involved in the determination of yeast Saccharomyces cerevisiae sensitivity to ionizing radiation].Genetika. 2008 Aug;44(8):1045-55. Genetika. 2008. PMID: 18825953 Russian.
-
RAD9, RAD17, and RAD24 are required for S phase regulation in Saccharomyces cerevisiae in response to DNA damage.Genetics. 1997 Jan;145(1):45-62. doi: 10.1093/genetics/145.1.45. Genetics. 1997. PMID: 9017389 Free PMC article.
-
Cell-cycle-specific activators of the Mec1/ATR checkpoint kinase.Biochem Soc Trans. 2011 Apr;39(2):600-5. doi: 10.1042/BST0390600. Biochem Soc Trans. 2011. PMID: 21428947 Review.
-
Dual cell cycle checkpoints sensitive to chromosome replication and DNA damage in the budding yeast Saccharomyces cerevisiae.Radiat Res. 1992 Nov;132(2):141-3. Radiat Res. 1992. PMID: 1438694 Review.
Cited by
-
Chromatin and the genome integrity network.Nat Rev Genet. 2013 Jan;14(1):62-75. doi: 10.1038/nrg3345. Nat Rev Genet. 2013. PMID: 23247436 Free PMC article. Review.
-
Novel connections between DNA replication, telomere homeostasis, and the DNA damage response revealed by a genome-wide screen for TEL1/ATM interactions in Saccharomyces cerevisiae.Genetics. 2013 Apr;193(4):1117-33. doi: 10.1534/genetics.113.149849. Epub 2013 Feb 1. Genetics. 2013. PMID: 23378069 Free PMC article.
-
The DNA damage checkpoint allows recombination between divergent DNA sequences in budding yeast.DNA Repair (Amst). 2011 Nov 10;10(11):1086-94. doi: 10.1016/j.dnarep.2011.07.007. Epub 2011 Oct 5. DNA Repair (Amst). 2011. PMID: 21978436 Free PMC article.
-
Site-specific phosphorylation of the DNA damage response mediator rad9 by cyclin-dependent kinases regulates activation of checkpoint kinase 1.PLoS Genet. 2013 Apr;9(4):e1003310. doi: 10.1371/journal.pgen.1003310. Epub 2013 Apr 4. PLoS Genet. 2013. PMID: 23593009 Free PMC article.
-
Stable hypermutators revealed by the genomic landscape of DNA repair genes among yeast species.bioRxiv [Preprint]. 2025 Mar 17:2025.03.15.643480. doi: 10.1101/2025.03.15.643480. bioRxiv. 2025. PMID: 40166188 Free PMC article. Preprint.
References
-
- Weinert TA, Hartwell LH. The RAD9 gene controls the cell cycle response to DNA damage in saccharomyces cerevisiae. Science. 1988;241:317–322. - PubMed
-
- Paulovich AG, Hartwell LH. A checkpoint regulates the rate of progression through S phase in S. cerevisiae in response to DNA damage. Cell. 1995;82:841–847. - PubMed
-
- Walsh T, King MC. Ten genes for inherited breast cancer. Cancer Cell. 2007;11:103–105. - PubMed
-
- Friedberg EC, Friedberg EC. DNA Repair and Mutagenesis. ASM Press; Washington, D.C: 2006.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

