A cell-free microtiter plate screen for improved [FeFe] hydrogenases
- PMID: 20479937
- PMCID: PMC2866662
- DOI: 10.1371/journal.pone.0010554
A cell-free microtiter plate screen for improved [FeFe] hydrogenases
Abstract
Background: [FeFe] hydrogenase enzymes catalyze the production and dissociation of H(2), a potential renewable fuel. Attempts to exploit these catalysts in engineered systems have been hindered by the biotechnologically inconvenient properties of the natural enzymes, including their extreme oxygen sensitivity. Directed evolution has been used to improve the characteristics of a range of natural catalysts, but has been largely unsuccessful for [FeFe] hydrogenases because of a lack of convenient screening platforms.
Methodology/principal findings: Here we describe an in vitro screening technology for oxygen-tolerant and highly active [FeFe] hydrogenases. Despite the complexity of the protocol, we demonstrate a level of reproducibility that allows moderately improved mutants to be isolated. We have used the platform to identify a mutant of the Chlamydomonas reinhardtii [FeFe] hydrogenase HydA1 with a specific activity approximately 4 times that of the wild-type enzyme.
Conclusions/significance: Our results demonstrate the feasibility of using the screen presented here for large-scale efforts to identify improved biocatalysts for energy applications. The system is based on our ability to activate these complex enzymes in E. coli cell extracts, which allows unhindered access to the protein maturation and assay environment.
Conflict of interest statement
Figures

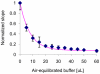
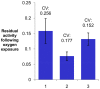

Similar articles
-
Synthetic biology for improved hydrogen production in Chlamydomonas reinhardtii.Microb Biotechnol. 2022 Jul;15(7):1946-1965. doi: 10.1111/1751-7915.14024. Epub 2022 Mar 26. Microb Biotechnol. 2022. PMID: 35338590 Free PMC article. Review.
-
Development of an in vitro compartmentalization screen for high-throughput directed evolution of [FeFe] hydrogenases.PLoS One. 2010 Dec 6;5(12):e15275. doi: 10.1371/journal.pone.0015275. PLoS One. 2010. PMID: 21151915 Free PMC article.
-
Genetic disruption of both Chlamydomonas reinhardtii [FeFe]-hydrogenases: Insight into the role of HYDA2 in H₂ production.Biochem Biophys Res Commun. 2012 Jan 13;417(2):704-9. doi: 10.1016/j.bbrc.2011.12.002. Epub 2011 Dec 8. Biochem Biophys Res Commun. 2012. PMID: 22177948
-
High-yield expression of heterologous [FeFe] hydrogenases in Escherichia coli.PLoS One. 2010 Nov 24;5(11):e15491. doi: 10.1371/journal.pone.0015491. PLoS One. 2010. PMID: 21124800 Free PMC article.
-
Approaches to developing biological H(2)-photoproducing organisms and processes.Biochem Soc Trans. 2005 Feb;33(Pt 1):70-2. doi: 10.1042/BST0330070. Biochem Soc Trans. 2005. PMID: 15667268 Review.
Cited by
-
A synthetic system links FeFe-hydrogenases to essential E. coli sulfur metabolism.J Biol Eng. 2011 May 26;5:7. doi: 10.1186/1754-1611-5-7. J Biol Eng. 2011. PMID: 21615937 Free PMC article.
-
Heterologous Hydrogenase Overproduction Systems for Biotechnology-An Overview.Int J Mol Sci. 2020 Aug 16;21(16):5890. doi: 10.3390/ijms21165890. Int J Mol Sci. 2020. PMID: 32824336 Free PMC article. Review.
-
Synthetic biology for improved hydrogen production in Chlamydomonas reinhardtii.Microb Biotechnol. 2022 Jul;15(7):1946-1965. doi: 10.1111/1751-7915.14024. Epub 2022 Mar 26. Microb Biotechnol. 2022. PMID: 35338590 Free PMC article. Review.
-
Second and Outer Coordination Sphere Effects in Nitrogenase, Hydrogenase, Formate Dehydrogenase, and CO Dehydrogenase.Chem Rev. 2022 Jul 27;122(14):11900-11973. doi: 10.1021/acs.chemrev.1c00914. Epub 2022 Jul 18. Chem Rev. 2022. PMID: 35849738 Free PMC article. Review.
-
Haplotype-Phased Synthetic Long Reads from Short-Read Sequencing.PLoS One. 2016 Jan 20;11(1):e0147229. doi: 10.1371/journal.pone.0147229. eCollection 2016. PLoS One. 2016. PMID: 26789840 Free PMC article.
References
-
- Adams MW. The structure and mechanism of iron-hydrogenases. Biochimica et Biophysica Acta. 1990;1020:115–145. - PubMed
-
- Mertens R, Liese A. Biotechnological applications of hydrogenases. Current Opinion in Biotechnology. 2004;15:343–348. - PubMed
-
- Levin D, Pitt L, Love M. Biohydrogen production: prospects and limitations to practical application. International Journal of Hydrogen Energy: 2004;29:173 –185.
-
- Prince RC, Kheshgi HS. The photobiological production of hydrogen: potential efficiency and effectiveness as a renewable fuel. Critical Reviews in Microbiology. 2005;31:19–31. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

