Functional characterization of alternate optimal solutions of Escherichia coli's transcriptional and translational machinery
- PMID: 20483314
- PMCID: PMC2872367
- DOI: 10.1016/j.bpj.2010.01.060
Functional characterization of alternate optimal solutions of Escherichia coli's transcriptional and translational machinery
Abstract
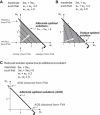
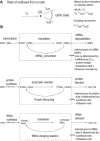
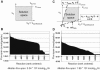
The constraint-based reconstruction and analysis approach has recently been extended to describe Escherichia coli's transcriptional and translational machinery. Here, we introduce the concept of reaction coupling to represent the dependency between protein synthesis and utilization. These coupling constraints lead to a significant contraction of the feasible set of steady-state fluxes. The subset of alternate optimal solutions (AOS) consistent with maximal ribosome production was calculated. The majority of transcriptional and translational reactions were active for all of these AOS, showing that the network has a low degree of redundancy. Furthermore, all calculated AOS contained the qualitative expression of at least 92% of the known essential genes. Principal component analysis of AOS demonstrated that energy currencies (ATP, GTP, and phosphate) dominate the network's capability to produce ribosomes. Additionally, we identified regulatory control points of the network, which include the transcription reactions of sigma70 (RpoD) as well as that of a degradosome component (Rne) and of tRNA charging (ValS). These reactions contribute significant variance among AOS. These results show that constraint-based modeling can be applied to gain insight into the systemic properties of E. coli's transcriptional and translational machinery.
Copyright 2010 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures



Similar articles
-
Genome-scale reconstruction of Escherichia coli's transcriptional and translational machinery: a knowledge base, its mathematical formulation, and its functional characterization.PLoS Comput Biol. 2009 Mar;5(3):e1000312. doi: 10.1371/journal.pcbi.1000312. Epub 2009 Mar 13. PLoS Comput Biol. 2009. PMID: 19282977 Free PMC article.
-
Preparation of Escherichia coli Rne protein and reconstitution of RNA degradosome.Methods Enzymol. 2001;342:346-56. doi: 10.1016/s0076-6879(01)42557-2. Methods Enzymol. 2001. PMID: 11586907 No abstract available.
-
Detachment of the RNA degradosome from the inner membrane of Escherichia coli results in a global slowdown of mRNA degradation, proteolysis of RNase E and increased turnover of ribosome-free transcripts.Mol Microbiol. 2019 Jun;111(6):1715-1731. doi: 10.1111/mmi.14248. Epub 2019 Apr 6. Mol Microbiol. 2019. PMID: 30903628 Free PMC article.
-
The Escherichia coli RNA degradosome: structure, function and relationship in other ribonucleolytic multienzyme complexes.Biochem Soc Trans. 2002 Apr;30(2):150-5. Biochem Soc Trans. 2002. PMID: 12035760 Review.
-
The RNA degradosome of Escherichia coli: an mRNA-degrading machine assembled on RNase E.Annu Rev Microbiol. 2007;61:71-87. doi: 10.1146/annurev.micro.61.080706.093440. Annu Rev Microbiol. 2007. PMID: 17447862 Review.
Cited by
-
Constraining the metabolic genotype-phenotype relationship using a phylogeny of in silico methods.Nat Rev Microbiol. 2012 Feb 27;10(4):291-305. doi: 10.1038/nrmicro2737. Nat Rev Microbiol. 2012. PMID: 22367118 Free PMC article. Review.
-
Enhanced flux prediction by integrating relative expression and relative metabolite abundance into thermodynamically consistent metabolic models.PLoS Comput Biol. 2019 May 13;15(5):e1007036. doi: 10.1371/journal.pcbi.1007036. eCollection 2019 May. PLoS Comput Biol. 2019. PMID: 31083653 Free PMC article.
-
Constraint-based models predict metabolic and associated cellular functions.Nat Rev Genet. 2014 Feb;15(2):107-20. doi: 10.1038/nrg3643. Epub 2014 Jan 16. Nat Rev Genet. 2014. PMID: 24430943 Review.
-
Genome-scale models of metabolism and gene expression extend and refine growth phenotype prediction.Mol Syst Biol. 2013 Oct 1;9:693. doi: 10.1038/msb.2013.52. Mol Syst Biol. 2013. PMID: 24084808 Free PMC article.
-
VFFVA: dynamic load balancing enables large-scale flux variability analysis.BMC Bioinformatics. 2020 Sep 29;21(1):424. doi: 10.1186/s12859-020-03711-2. BMC Bioinformatics. 2020. PMID: 32993482 Free PMC article.
References
-
- Laffend L., Shuler M.L. Ribosomal protein limitations in Escherichia coli under conditions of high translational activity. Biotechnol. Bioeng. 1994;43:388–398. - PubMed
-
- Bremer H., Dennis P., Ehrenberg M. Free RNA polymerase and modeling global transcription in Escherichia coli. Biochimie. 2003;85:597–609. - PubMed
-
- Alberghina L., Mariani L. Analysis of a cell cycle model for Escherichia coli. J. Math. Biol. 1980;9:389–398. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

