African-derived genetic polymorphisms in TNFAIP3 mediate risk for autoimmunity
- PMID: 20483768
- PMCID: PMC3307531
- DOI: 10.4049/jimmunol.1000324
African-derived genetic polymorphisms in TNFAIP3 mediate risk for autoimmunity
Abstract
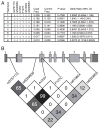
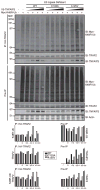
The TNF alpha-induced protein 3 (TNFAIP3) is an ubiquitin-modifying enzyme and an essential negative regulator of inflammation. Genome-wide association studies have implicated the TNFAIP3 locus in susceptibility to autoimmune disorders in European cohorts, including rheumatoid arthritis, coronary artery disease, psoriasis, celiac disease, type 1 diabetes, inflammatory bowel disease, and systemic lupus erythematosus (SLE). There are two nonsynonymous coding polymorphisms in the deubiquitinating (DUB) domain of TNFAIP3: F127C, which is in high-linkage disequilibrium with reported SLE-risk variants, and A125V, which has not been previously studied. We conducted a case-control study in African-American SLE patients using these coding variants, along with tagging polymorphisms in TNFAIP3, and identified a novel African-derived risk haplotype that is distinct from previously reported risk variants (odds ratio=1.6, p=0.006). In addition, a rare protective haplotype was defined by A125V (odds ratio=0.31, p=0.027). Although A125V was associated with protection from SLE, surprisingly the same allele was associated with increased risk of inflammatory bowel disease. We tested the functional activity of nonsynonymous coding polymorphisms within TNFAIP3, and found that the A125V coding-change variant alters the DUB activity of the protein. Finally, we used computer modeling to depict how the A125V amino acid change in TNFAIP3 may affect the three-dimensional structure of the DUB domain to a greater extent than F127C. This is the first report of an association between TNFAIP3 polymorphisms and autoimmunity in African-Americans.
Conflict of interest statement
The authors have no financial conflicts of interest.
Figures


References
-
- Wertz IE, O’Rourke KM, Zhou H, Eby M, Aravind L, Seshagiri S, Wu P, Wiesmann C, Baker R, Boone DL, et al. De-ubiquitination and ubiquitin ligase domains of A20 downregulate NF-kappaB signalling. Nature. 2004;430:694–699. - PubMed
-
- Boone DL, Turer EE, Lee EG, Ahmad RC, Wheeler MT, Tsui C, Hurley P, Chien M, Chai S, Hitotsumatsu O, et al. The ubiquitin-modifying enzyme A20 is required for termination of Toll-like receptor responses. Nat Immunol. 2004;5:1052–1060. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 1F32AI077235-01A1/AI/NIAID NIH HHS/United States
- P30 DK042086/DK/NIDDK NIH HHS/United States
- F32 AI077235/AI/NIAID NIH HHS/United States
- UL1 RR024999/RR/NCRR NIH HHS/United States
- U01 CA153060/CA/NCI NIH HHS/United States
- L30 AI071651/AI/NIAID NIH HHS/United States
- K08AI083790/AI/NIAID NIH HHS/United States
- T32AI-07090/AI/NIAID NIH HHS/United States
- K08 AI083790/AI/NIAID NIH HHS/United States
- T32 DK007074/DK/NIDDK NIH HHS/United States
- AI071651/AI/NIAID NIH HHS/United States
- DK07074/DK/NIDDK NIH HHS/United States
- T32 AI007090/AI/NIAID NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

