Divinyl chlorophyll(ide) a can be converted to monovinyl chlorophyll(ide) a by a divinyl reductase in rice
- PMID: 20484022
- PMCID: PMC2899930
- DOI: 10.1104/pp.110.158477
Divinyl chlorophyll(ide) a can be converted to monovinyl chlorophyll(ide) a by a divinyl reductase in rice
Abstract
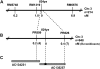
3,8-Divinyl (proto)chlorophyll(ide) a 8-vinyl reductase (DVR) catalyzes the reduction of 8-vinyl group on the tetrapyrrole to an ethyl group, which is indispensable for monovinyl chlorophyll (Chl) synthesis. So far, three 8-vinyl reductase genes (DVR, bciA, and slr1923) have been characterized from Arabidopsis (Arabidopsis thaliana), Chlorobium tepidum, and Synechocystis sp. PCC6803. However, no 8-vinyl reductase gene has yet been identified in monocotyledonous plants. In this study, we isolated a spontaneous mutant, 824ys, in rice (Oryza sativa). The mutant exhibited a yellow-green leaf phenotype, reduced Chl level, arrested chloroplast development, and retarded growth rate. The phenotype of the 824ys mutant was caused by a recessive mutation in a nuclear gene on the short arm of rice chromosome 3. Map-based cloning of this mutant resulted in the identification of a gene (Os03g22780) showing sequence similarity with the Arabidopsis DVR gene (AT5G18660). In the 824ys mutant, nine nucleotides were deleted at residues 952 to 960 in the open reading frame, resulting in a deletion of three amino acid residues in the encoded product. High-performance liquid chromatography analysis of Chls indicated the mutant accumulates only divinyl Chl a and b. A recombinant protein encoded by Os03g22780 was expressed in Escherichia coli and found to catalyze the conversion of divinyl chlorophyll(ide) a to monovinyl chlorophyll(ide) a. Therefore, it has been confirmed that Os03g22780, renamed as OsDVR, encodes a functional DVR in rice. Based upon these results, we succeeded to identify an 8-vinyl reductase gene in monocotyledonous plants and, more importantly, confirmed the DVR activity to convert divinyl Chl a to monovinyl Chl a.
Figures





References
-
- Adra AN, Rebeiz CA. (1998) Chloroplast biogenesis 81: transient formation of divinyl chlorophyll a following a 2.5 ms light flash treatment of etiolated cucumber cotyledons. Photochem Photobiol 68: 852–856
-
- Bazzaz MB. (1981) New chlorophyll chromophores isolate from a chlorophyll deficient mutant of maize. Photobiochem Photobiophys 2: 199–207
-
- Bazzaz MB, Bradley CV, Brereton RG. (1982) 4-Vinyl-4-desethyl chlorophyll a: characterization of a new naturally occurring chlorophyll using fast atom bombardment, field desorption and “in beam” electron impact mass spectroscopy. Tetrahedron Lett 23: 1211–1214
-
- Bazzaz MB, Brereton RG. (1982) 4-Vinyl-4-desethyl chlorophyll a: a new naturally occurring chlorophyll. FEBS Lett 138: 104–108
-
- Beale SI. (1999) Enzymes of chlorophyll biosynthesis. Photosynth Res 60: 47–73
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

