Genome-wide DNA methylation profiling of chronic lymphocytic leukemia allows identification of epigenetically repressed molecular pathways with clinical impact
- PMID: 20484983
- PMCID: PMC3322493
- DOI: 10.4161/epi.5.6.12179
Genome-wide DNA methylation profiling of chronic lymphocytic leukemia allows identification of epigenetically repressed molecular pathways with clinical impact
Abstract
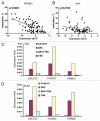
We performed a genome-wide analysis of aberrant DNA methylation in chronic lymphocytic leukemia (CLL) using methylated CpG island amplification (MCA) coupled with a promoter microarray. We identified 280 potential targets of aberrant DNA methylation in CLL. These genes were located more frequently in chromosomes 19 (16%, p=0.001), 16 (11%, p=0.001), 17 (10%, p=0.02) and 11 (9%, p=0.02) and could be grouped in several functional networks. Methylation status was confirmed for 22 of these genes (SOX11, DLX1, FAM62C, SOX14, RSPO1, ADCY5, HAND2,SPOCK, MLL, ING1, PRIMA1, BCL11B, LTBP2, BNC1, NR2F2, SALL1, GALGT2, LHX1, DLX4, KLK10, TFAP2 and APP) in 78 CLL patients by pyrosequencing. As a proof of principle, we analyzed the expression of 2 genes, PRIMA1 and APP, in primary cells and of GALGT2, TFAP2C and PRIMA1 in leukemia cells. There was an inverse association between methylation and gene expression. This could be reversed by treatment with 5-aza-2'-deoxycytidine in cell lines. Treatment in a clinical trial with 5-azacitidine resulted in decreased methylation of LINE, DLX4 and SALL1 in the peripheral blood B-cells of patients with CLL. IgVH mutational status or ZAP-70 expression were not associated with specific methylation profiles. By multivariate analysis, methylation of LINE and APP was associated with shorter overall survival (p = 0.045 and 0.0035, respectively). This study demonstrates that aberrant DNA methylation is common and has potential prognostic and therapeutic value in CLL.
Figures



References
-
- Chiorazzi N, Rai KR, Ferrarini M. Chronic lymphocytic leukemia. New Eng J Med. 2005;352:804–815. - PubMed
-
- Keating MJ, Chiorazzi N, Messmer B, Damle RN, Allen SL, Rai KR, et al. Biology and treatment of chronic lymphocytic leukemia. Hematology Am Soc Hematol Educ Program. 2003:153–175. - PubMed
-
- Rai KR, Peterson BL, Appelbaum FR, Kolitz J, Elias L, Shepherd L, et al. Fludarabine compared with chlorambucil as primary therapy for chronic lymphocytic leukemia. New Eng J Med. 2000;343:1750–1757. - PubMed
-
- O'Brien SM, Kantarjian H, Thomas DA, Giles FJ, Freireich EJ, Cortes J, et al. Rituximab dose-escalation trial in chronic lymphocytic leukemia. J Clin Oncol. 2001;19:2165–2170. - PubMed
-
- Byrd JC, Rai K, Peterson BL, Appelbaum FR, Morrison VA, Kolitz JE, et al. Addition of rituximab to fludarabine may prolong progression-free survival and overall survival in patients with previously untreated chronic lymphocytic leukemia: an updated retrospective comparative analysis of CALGB 9712 and CALGB 9011. Blood. 2005;105:49–53. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
