Arabidopsis synaptotagmin SYT1, a type I signal-anchor protein, requires tandem C2 domains for delivery to the plasma membrane
- PMID: 20498364
- PMCID: PMC2906310
- DOI: 10.1074/jbc.M109.084046
Arabidopsis synaptotagmin SYT1, a type I signal-anchor protein, requires tandem C2 domains for delivery to the plasma membrane
Abstract
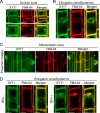
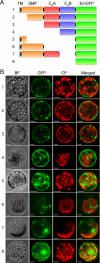
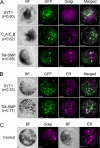
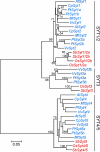
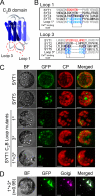
The correct localization of integral membrane proteins to subcellular compartments is important for their functions. Synaptotagmin contains a single transmembrane domain that functions as a type I signal-anchor sequence in its N terminus and two calcium-binding domains (C(2)A and C(2)B) in its C terminus. Here, we demonstrate that the localization of an Arabidopsis synaptotagmin homolog, SYT1, to the plasma membrane (PM) is modulated by tandem C2 domains. An analysis of the roots of a transformant-expressing green fluorescent protein-tagged SYT1 driven by native SYT1 promoter suggested that SYT1 is synthesized in the endoplasmic reticulum, and then delivered to the PM via the exocytotic pathway. We transiently expressed a series of truncated proteins in protoplasts, and determined that tandem C(2)A-C(2)B domains were necessary for the localization of SYT1 to the PM. The PM localization of SYT1 was greatly reduced following mutation of the calcium-binding motifs of the C(2)B domain, based on sequence comparisons with other homologs, such as endomembrane-localized SYT5. The localization of SYT1 to the PM may have been required for the functional divergence that occurred in the molecular evolution of plant synaptotagmins.
Figures


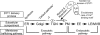
Similar articles
-
Arabidopsis SYT1 maintains stability of cortical endoplasmic reticulum networks and VAP27-1-enriched endoplasmic reticulum-plasma membrane contact sites.J Exp Bot. 2016 Nov;67(21):6161-6171. doi: 10.1093/jxb/erw381. Epub 2016 Oct 17. J Exp Bot. 2016. PMID: 27811083 Free PMC article.
-
The Arabidopsis synaptotagmin1 is enriched in endoplasmic reticulum-plasma membrane contact sites and confers cellular resistance to mechanical stresses.Plant Physiol. 2015 May;168(1):132-43. doi: 10.1104/pp.15.00260. Epub 2015 Mar 19. Plant Physiol. 2015. PMID: 25792253 Free PMC article.
-
Synaptotagmin-Associated Endoplasmic Reticulum-Plasma Membrane Contact Sites Are Localized to Immobile ER Tubules.Plant Physiol. 2018 Oct;178(2):641-653. doi: 10.1104/pp.18.00498. Epub 2018 Aug 20. Plant Physiol. 2018. PMID: 30126867 Free PMC article.
-
Biological roles of plant synaptotagmins.Eur J Cell Biol. 2023 Sep;102(3):151335. doi: 10.1016/j.ejcb.2023.151335. Epub 2023 Jun 20. Eur J Cell Biol. 2023. PMID: 37390668 Review.
-
Models of synaptotagmin-1 to trigger Ca2+ -dependent vesicle fusion.FEBS Lett. 2018 Nov;592(21):3480-3492. doi: 10.1002/1873-3468.13193. Epub 2018 Jul 30. FEBS Lett. 2018. PMID: 30004579 Review.
Cited by
-
Geometry and cellular function of organelle membrane interfaces.Plant Physiol. 2021 Apr 2;185(3):650-662. doi: 10.1093/plphys/kiaa079. Plant Physiol. 2021. PMID: 33793898 Free PMC article. Review.
-
Arabidopsis SYT1 maintains stability of cortical endoplasmic reticulum networks and VAP27-1-enriched endoplasmic reticulum-plasma membrane contact sites.J Exp Bot. 2016 Nov;67(21):6161-6171. doi: 10.1093/jxb/erw381. Epub 2016 Oct 17. J Exp Bot. 2016. PMID: 27811083 Free PMC article.
-
D6 protein kinase in root xylem benefiting resistance to Fusarium reveals infection and defense mechanisms in tung trees.Hortic Res. 2021 Nov 1;8(1):240. doi: 10.1038/s41438-021-00656-2. Hortic Res. 2021. PMID: 34719680 Free PMC article.
-
Rare earth elements induce cytoskeleton-dependent and PI4P-associated rearrangement of SYT1/SYT5 endoplasmic reticulum-plasma membrane contact site complexes in Arabidopsis.J Exp Bot. 2020 Jul 6;71(14):3986-3998. doi: 10.1093/jxb/eraa138. J Exp Bot. 2020. PMID: 32179893 Free PMC article.
-
Plasma membrane localization of Solanum tuberosum remorin from group 1, homolog 3 is mediated by conformational changes in a novel C-terminal anchor and required for the restriction of potato virus X movement].Plant Physiol. 2012 Oct;160(2):624-37. doi: 10.1104/pp.112.200519. Epub 2012 Aug 1. Plant Physiol. 2012. PMID: 22855937 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

